Search
Your Data: Isolating miRNAs from Pancreatic Samples
Introduction
Recently, 19–23 nt RNA molecules, known as microRNAs or miRNAs, have been shown to be critical for gene expression regulation. As many as several thousand miRNAs have already been identified and are predicted to regulate 30–50% of protein-encoding genes. Isolation and expression profiling of these molecules has become important to identify miRNAs involved in specific cellular processes and diseases, and to link them to potential target gene function. However, standard RNA isolation methods do not always retain the small RNA fraction.
Ambion specializes in developing tools for RNA analysis, including RNA isolation. The mirVana™ miRNA Isolation Kit purifies total RNA that includes quantitative amounts of small RNAs. It also includes a protocol for enrichment of the small RNA fraction. Drs. Sipos and Hahn used this kit to isolate total RNA from pancreatic samples for subsequent analysis of miRNA expression.
Drs. Bence Sipos and Stephan Hahn
Ruhr-University Bochum, Molecular GI-Oncology (MGO), Department of Internal Medicine, UniversitätsstraΒe 150, 44780 Bochum, Germany. Tel: +49 234 3229282; Fax: +49 234 3214674; Email: stephan.hahn@rub.de and Department of Pathology, University of Kiel, Michaelistr. 11, 24105 Kiel, Tel +49-431-597-3402
Involvement of miRNAs in Pancreatic Disease
Dr. Hahn and colleagues study pancreatic cancer development, especially the identification of new diagnostic and therapeutic targets for this disease. The search for dysregulated miRNAs in pancreatic carcinoma tissues opens a new avenue for identifying such targets.
Given that the expression of specific miRNAs has been linked with several diseases [1−5], the research group wanted to address the potential involvement of miRNA in pancreatic cancer. The first step in this study was to isolate good quality RNA from normal and diseased pancreatic tissues using an efficient RNA isolation method that would retain the miRNA fraction.
Preparing Samples for miRNA Analyses
Pancreatic tissue is known to contain extremely high levels of endogenous RNase activity making isolation of intact RNA challenging. However, the tumor and chronic pancreatitis samples used in this study contained very few RNase-producing acinar cells, and consequently, had much lower RNase content compared to normal pancreatic tissue. To prevent RNA degradation by endogenous RNases, the normal pancreas specimens were saturated with RNAlater® Tissue Collection:RNA Stabilization Solution (Ambion) prior to long term storage at −80°C. The tumor and chronic pancreatitis specimens were directly flash frozen and stored at −80°C.
Total RNA was isolated from all samples using the mirVana™ miRNA Isolation Kit and was analyzed on a denaturing gel (Figure 1). RNA from the majority of the processed samples was of very good quality.
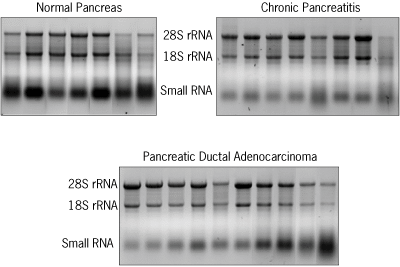
Figure 1. Total RNA Isolated from Normal and Diseased Pancreas Samples. Total RNA, including the small RNA fraction, was isolated from 25 pancreatic samples from donors with either normal tissue, pancreatic carcinoma, or chronic pancreatitis as indicated, using the mirVana™ miRNA Isolation Kit. RNA (3 µg) was analyzed on a 1% formaldehyde agarose gel.
The mirVana miRNA Isolation Kit also allows for separation of RNA species smaller than 200 nt (miRNAs, tRNAs, 5S and 5.8S rRNAs, snRNAs) from long RNA species (18S and 28S rRNAs, mRNAs). Fractions enriched in small RNAs can be used in downstream miRNA expression analyses, and are particularly useful for Northern blotting and solution hybridization assays (RPA). For miRNA array analysis applications, we recommend isolating total RNA (including the small RNA fraction), and enriching for the small RNA fraction using Ambion’s flashPAGE™ Fractionator to obtain a significant increase in sensitivity compared to total RNA samples, as well as to separate the mature miRNA population from the pre-miRNA population. Finally, for real-time PCR applications, such as for use with Applied Biosystems TaqMan® MicroRNA Assays, total RNA is recommended.
Figure 2 demonstrates the quality of total RNA and enriched small RNA fractions obtained from 5 different pancreatic cell lines using the mirVana miRNA Isolation Kit. Note the complete removal of the large, abundant rRNA species from the small RNA enriched fraction. The RNA samples from these two experiments are currently being used for miRNA expression profiling using mirVana miRNA Bioarrays and mirVana miRNA qRT-PCR Kits with the goal of identifying key miRNAs involved in pancreatic carcinogenesis.
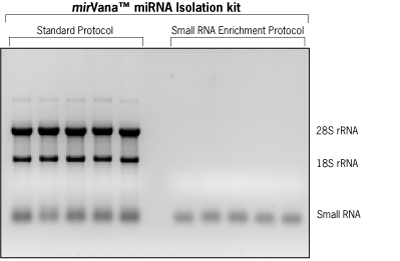
Figure 2. Total RNA and RNA Enriched for the Small RNA Fraction Isolated from Cultured Pancreatic Cell Lines. RNA was isolated from 5 different pancreatic cell lines, using either the standard mirVana™ miRNA Isolation Kit protocol, or the small RNA enrichment protocol. Total RNA (3 µg) and fractions enriched in small RNAs (300 ng) from each of the samples were analyzed on a 1% formaldehyde containing agarose gel.
mirVana™ miRNA Isolation Kit
The mirVana miRNA Isolation Kit efficiently isolates small RNA-containing total RNA from 40 samples of 102–107 cultured cells or from 0.5–250 mg tissue in a simple 30 minute procedure. In addition, it can be used to enrich for small RNA <200 nt. Not limited to miRNA, the procedure is ideal for isolating all small RNA species, including siRNA, shRNA, and snRNAs, and is compatible with virtually all cell and tissue types.
mirVana™ miRNA Bioarrays are high quality microarrays spotted with probes targeting a comprehensive selection of human, mouse, and rat miRNAs from Version 8.0 of the miRBase Sequence database. In addition, mirVana miRNA Bioarrays include probes targeting an exclusive set of newly identified human miRNAs. Called Ambi-miRs™, these novel miRNAs were identified through a combination of bioinformatic prediction, cloning, and detection in human total RNA samples. For researchers who prefer to prepare their own microarrays, the mirVana miRNA Probe Set is available. It contains the same content as the mirVana miRNA Bioarrays. Both products are updated to include newly identified miRNAs in the miRBase Sequence database.
Applied Biosystems TaqMan® MicroRNA Assays quantitate miRNAs with the specificity and sensitivity of gold standard TaqMan Assay chemistry. Choose from hundreds of new, individual real-time PCR assays for human, mouse, and rat. This new assay collection offers the ultimate flexibility in your miRNA research, with assays for C. elegans, Arabidopsis, and Drosophila coming soon.
- Quantitate only biologically active, mature miRNAs
- Requires only 1–10 ng of total RNA or equivalent
- Up to seven logs of dynamic range
- High-quality results in under three hours
- Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, Croce CM (2004) Human miRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101:2999−3004.
- Metzler M, Wilda M, Busch K, Viehmann S, Borkhardt A (2004) High expression of precursor miRNA-155/BIC RNA in children with Burkitt lymphoma. Genes Chrom Cancer 2:167−9.
- Michael MZ, O’Connor SM, van Holst Pellekaan NG, Young GP, James RJ (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Molec Cancer Res 1:882−91.
- Esquela-Kerscher A, Slack FJ (2006) Oncomirs–microRNAs with a role in cancer. Nat Rev Cancer 6(4):259−69.
- Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A, Zlotorynski E, Yabuta N, De Vita G, Nojima H, Looijenga LH, Agami R (2006) A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 124(6):1169−81