Search
A Better Way to Isolate RNA AND Protein From the Same Sample
- Isolate total RNA and protein from cultured cells or tissues
- Isolate nuclear and cytoplasmic RNA and protein from cultured cells
- No phenol extraction or alcohol precipitation
Our new PARIS™ Kit uses a fast, simple procedure based on our RNAqueous® technology to isolate both RNA and native protein from the same experimental sample. The kit also permits separation of nuclear and cytoplasmic fractions prior to RNA and/or protein isolation. The resulting protein and RNA samples are suitable for various downstream applications, reducing time, cost, and variability between independent experimental samples.
Protein And RNA Isolation System (PARIS)
Isolation of high quality RNA is the first step for a variety of gene expression analyses. Very often complementary studies at the protein level are also required, e.g. performing further in vitro functional assays, analyzing a regulatory pathway, or simply correlating RNA and protein expression levels. Usually these analyses are performed using different aliquots of the same experimental sample. However, when working with rare, difficult to obtain, or very small samples it is sometimes impractical to isolate RNA and native proteins independently. In studies involving large numbers of samples, expensive reagents, or inherent variability (e.g. cell transfection), the addition of independent experimental samples is not only costly and time consuming, but may also lead to inconsistent results. To solve these issues, our scientists have developed a unique Protein And RNA Isolation System (PARIS) that allows researchers to isolate both RNA and protein from the same experimental sample (Figure 1).
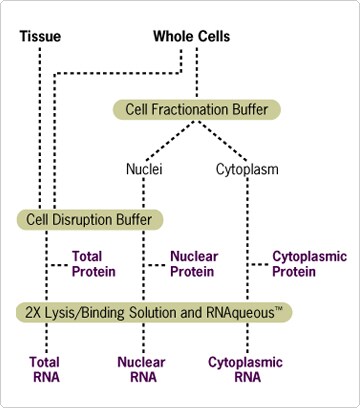
Figure 1. Schematic of PARIS Procedure.
Using PARIS, RNA and protein can be isolated simultaneously from whole cell lysates. Alternatively, RNA and protein can be isolated from separate nuclear and cytoplasmic fractions (see "Application: Nuclear vs. Cytoplasmic Fractionation" at right). Tissue or cultured cells are first homogenized in ice-cold Cell Disruption Buffer to prepare a total cell lysate. Since the homogenization is performed quickly on ice and in the presence of detergent, both protein and RNA can be purified directly from this lysate. For RNA isolation, a part of the total cell lysate is immediately mixed with an equal volume of Lysis/Binding Solution. This solution contains a high concentration of guanidinium thiocyanate, a strong chaotropic denaturant that rapidly inactivates cellular ribonucleases. Total RNA is then purified from the mixture using an RNA binding glass fiber filter. After three rapid washing steps, high quality RNA is eluted in a concentrated form. The entire procedure can be completed in less than 20 minutes. Note: This kit is not recommended for tissues with high levels of ribonucleases, such as pancreas.
Compatible with Most Downstream Applications
The RNA isolated from total, nuclear, or cytoplasmic fractions with the PARIS procedure can be used in a variety of downstream applications, including blot hybridization, in vitro translation, cDNA synthesis, and RT-PCR. A DNase I treatment is recommended for RNA that will be used for RT-PCR experiments, especially if using primers that do not flank introns, or for genes that have processed pseudogenes. Our DNA-free™ DNase Treatment and Removal Reagents (available separately) are ideal to quickly remove trace amounts of DNA from the total and nuclear RNA samples without phenol extraction or alcohol precipitation. The cytoplasmic RNA fraction is virtually free of DNA contamination.
Each protein fraction can also be used directly for most common applications, including functional assays, immunoprecipitation, Western blotting or two-dimensional gel electrophoresis (Figure 2 below). For experiments such as gel shift assays or enzymatic assays, protein samples are usually concentrated enough to be diluted in the final reaction buffer. The Cell Disruption Buffer and Cell Fractionation Buffer have been specially designed to be compatible with downstream analyses.
The PARIS Kit includes sufficient reagent to perform 50 purifications. The kit also comes with a comprehensive Instruction Manual and RNase-free microfuge tubes, glass fiber filters and reagents for RNA isolation. Each purification can accommodate 1 to 75 mg of tissue or 100 to 107 cells.
PARIS is Compatible with RNAlater® and RNAlater-ICE Treated Tissue.
Many researchers use the RNA stabilization reagent RNAlater to protect RNA integrity immediately after cell or tissue collection, to stabilize RNA during long period of storage, or to circumvent the need to grind frozen tissue into a powder prior to RNA isolation. (See the article "RNArticles: Keeping RNA Intact Prior to RNA Isolation - RNAlater®" for some of its many applications.) Our new RNAlater-ICE permits transition of previously frozen samples to a state compatible with handling at room temperature and standard homogenization methods. As shown in Figure 3, the PARIS procedure is compatible with both of these tissue collection/RNA stabilization reagents for both cells and tissue. After homogenization, protein samples are readily available for applications that do not require native functional protein, such as Western blot analysis.

Figure 2. 2D Analysis of Proteins Isolated Using the PARIS Kit. Protein samples were resolved using a pH 4-7 IPG gel followed by 8-16% SDS-PAGE. Top: Total protein (~25 µg) from HeLa cells stained with SYPRO® Ruby. Bottom: Total protein (~125 µg) from mouse brain stained with Coomassie Blue.
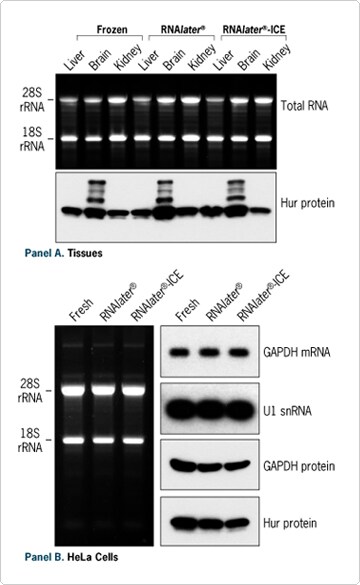
Figure 3. RNA and Protein Isolation from Stabilized Samples. (A) Isolation from ~30 mg of mouse liver, brain or kidney 5 days after collection. Samples were either kept frozen at -80ºC, stored in RNAlater® at 4ºC, or snap frozen and then stored in RNAlater-ICE at -20ºC. (B) Isolation from 106 HeLa cells freshly harvested, kept 2 days in RNAlater at RT, or in RNAlater-ICE at -20°C.
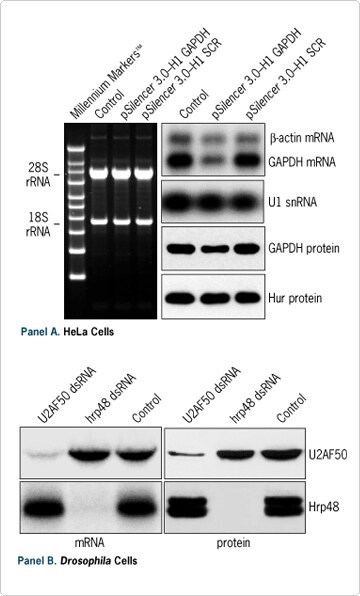
Figure 4. Analysis of the RNAi Effect. (A) 5 x 104 HeLa cells were transfected in a 24 well plate with the indicated plasmids. 48 hours after transfection, total RNA and protein were isolated with the PARIS Kit and analyzed by denaturing agarose gel electrophoresis, Northern blot, or Western blot. SCR represents a scrambled siRNA sequence. (B) 5 x 10 5 Schneider's Drosophila cells were treated for 48 hours in a 6 well plate with 10 nM of the indicated dsRNA prepared with our MEGAscript® RNAi Kit. Total RNA and protein were isolated with the PARIS Kit and analyzed by Northern blot and Western blot.
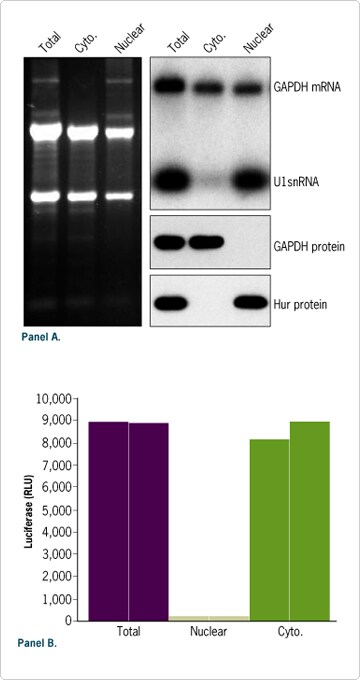
Figure 5. Efficient Cellular Fractionation. (A) RNA and protein were isolated from 106 HeLa cells using the PARIS Kit. To allow direct comparison between RNA isolated from different cellular compartments, the same volume of each fraction (~5%) was analyzed by denaturing agarose gel electrophoresis, Northern blot, or Western blot. (B) 3 x 10 5 HeLa cells were transfected in duplicate in 6 well plates with 1 µg of luciferase mRNA and the relative luciferase activity in each protein fraction was measure 18 hours after transfection. Luciferase mRNA was prepared with our mMESSAGE mMACHINE® and Poly(A) Tailing Kits.
- Analysis of RNA and Protein Expression After RNAi
Researchers often need to assess the specificity of their gene silencing experiments and to correlate knockdown of a target mRNA with the corresponding protein levels. Figure 4 demonstrates the effectiveness of the PARIS Kit for preparing samples for analyzing the effect of RNAi at both the RNA and protein levels in both mammalian and insect cells.
Here a specific reduction of GAPDH expression was observed both at the RNA level (~65%) and protein level (~50%) in HeLa cells transfected with a p Silencer 3.0-H1 Vector expressing an siRNA targeting the GAPDH gene (see Figure 4A). In contrast, no reduction was detected in levels of control RNA or proteins, nor was expression affected by transfection of a control plasmid expressing a scrambled siRNA sequence. Another example of RNAi analysis is shown in Figure 4B.
- Nuclear vs. Cytoplasmic Fractionation
Scientists studying nucleocytoplasmic transport, splicing, transcription, RNA stability, translation, or signal transduction may be interested in the nuclear vs. cytoplasmic distribution of various proteins or RNA. PARIS is the only fully integrated system that allows isolation of RNA and/or protein from these cellular fractions. For this purpose, the PARIS Kit contains a solution optimized for rapid nuclear/cytoplasmic fractionation of fresh cultured cells to be used prior to the protein/RNA isolation protocols described above. The fractionation method is based on differential lysis of the plasma and nuclear membranes by nonionic detergents. After selective lysis of the plasma membrane in Cell Fractionation Buffer, intact nuclei are collected by a quick centrifugation step and resuspended in Cell Disruption Buffer (see Figure 1). The supernatant contains all of the cytoplasmic components. Following the same procedure described above, both protein and high quality RNA can then be isolated from each cellular
fraction. The entire procedure can be completed in 30 minutes.
Efficient RNA and protein fractionation with the PARIS procedure is illustrated in Figure 5A. As expected, nuclear specific components such as precursor ribosomal RNA, U1 snRNA, and Hur protein were detected only in the nuclear fraction. In contrast, GAPDH protein was detected only in the cytoplasmic fraction. Efficient cellular fractionation was also observed after transfection of a firefly luciferase reporter mRNA in HeLa cells (Figure 5B). For each experiment total RNA and protein fractions prepared with the PARIS Kit were also analyzed. The sum of the signal generated from nuclear and cytoplasmic fractions was equivalent to the signal detected in the total control sample, confirming efficient fractionation and complete recovery of RNA and protein from each cellular fraction.