Search
High-Resolution Melt Analysis for Mutation Scanning Prior to Sequencing
Download the PDF version of this web page: High-Resolution Melt Analysis for Mutation Scanning Prior to Sequencing
Analyzing Genetic Variation Using HRM Mutation Scanning Followed by Sanger Sequencing - Making it Easier to Unlock Understanding
Mutation Scanning to Identify Candidates for Sequencing
Detailed characterization of genetic variation is becoming increasingly important for the understanding and treatment of genetic diseases and conditions. It is also useful in the development of biomarkers for use in drug development and individualized medicine. Sanger DNA sequencing remains the most straightforward, reliable method for determining the precise genetic sequence of DNA fragments; however, it requires a fairly significant investment in time and materials. Mutation scanning strategies seek to quickly and efficiently scan DNA samples from many individuals for minor genetic variations to identify candidates to be subjected to full sequencing analysis.

Figure 1. Overview of Workflow for Analysis of Genetic Variation With HRM Pre-screening.
HRM Analysis for Quicker, Easier, and More Reliable Mutation Scanning
With the introduction of brighter DNA-binding dyes, real-time PCR instruments that collect fluorescence data at finer temperature resolution, and intuitive software platforms, high-resolution melt (HRM) analysis is becoming the method of choice for scanning large numbers of samples for genetic variants. The procedure consists of amplifying the gene of interest in 80–250 bp segments in reactions that contain a fluorescent dsDNA-binding dye.
After PCR, the annealed reaction products are gradually denatured (or melted), and the resulting decrease in fluorescence as the DNA becomes single-stranded is measured in real time. The shape of the high-resolution melt curve generated will vary with different DNA sequences, providing data capable of distinguishing amplicons that differ by as little as a single base pair. HRM analysis can be used for detection of minor genetic variations such as single nucleotide polymorphisms, insertions, deletions, and inversions. For a thorough introduction to HRM analysis, including information on experimental design, optimization, and technical problem solving, download the Applied Biosystems Guide to HRM Analysis at: www.appliedbiosystems.com/hrm.
Compared to other technologies for mutation scanning such as sequencing, denaturing HPLC (dHPLC), and denaturing gradient gel electrophoresis (DGGE), HRM offers a far easier, less time-consuming procedure that consumes significantly less reagent. It is more reproducible than either dHPLC or DGGE, requiring less optimization and interpretation by the investigator. In some cases, it also provides better accuracy and sensitivity. In addition, HRM analysis is a closed-tube procedure, greatly reducing the risk of contamination from earlier PCR products. Because HRM analysis does not destroy samples, it is particularly well suited for genetic variant scanning. The HRM process identifies samples for further study, and the PCR reaction products can then be used as templates for DNA sequencing with minimal downstream processing. This application note outlines this workflow: detection of genomic variation using HRM mutation scanning followed by Sanger sequencing analysis (Figure 1). The procedure employs Applied Biosystems® reagents for HRM screening that not only provide superior performance compared to other commercially available dyes and enzyme mixtures, but have been extensively tested and optimized for HRM screening on the Applied Biosystems® HRM-capable StepOne™, StepOnePlus™, 7500 Fast, 7900HT Fast, and ViiA™ 7 Real-Time PCR Systems. Data are analyzed using Applied Biosystems® High Resolution Melt Software to identify samples for precise DNA sequence analysis using our industry-leading DNA sequencing reagents and capillary electrophoresis systems.
Step 1: PCR Primer Design
A critical first step in setting up mutation scanning experiments using HRM analysis is to design PCR primers to amplify overlapping segments of the genomic region of interest. Important considerations for the primer design step include designing appropriately sized amplicons and observing well-established conventions for designing high-quality PCR primers.
Select Overlapping 80–250 bp Target Regions
HRM analysis works well with amplicons in the ~120–300 bp size range; this size corresponds to 80–250 bp genomic DNA targets amplified using ~20–40 nt PCR primers. There is a balance between keeping amplicons short enough to yield simple, easily analyzed melt profiles, yet long enough to minimize the number of individual amplicons needed to cover the sequence. Large sequence variations may be detectable in amplicons longer than ~300 bp; however, more subtle variations such as A/T class 4 SNPs are typically detectable only in smaller amplicons. Since amplified PCR fragments that exhibit genetic variation will be further analyzed by Sanger DNA sequencing, you may want to include M13 universal sequencing primer tails on the 5’ end of each PCR primer. This significantly reduces the number of pipetting steps in the DNA sequencing portion of the workflow because, with M13 tails, all of the samples can be sequenced using generic M13 sequencing primers, rather than requiring specific primers for each amplicon. The extra M13 sequence does not affect the results of HRM analysis; however, it does change the shape of the melt profile somewhat, as would any change to the sequence of the amplicon (Figure 2). The drawback to including M13 sequence in the PCR primers is that it adds 36 bp (18 nt/PCR primer) to amplicon lengths, thereby decreasing the amount of genomic sequence queried in each amplicon (potentially increasing the number of amplicons required to cover the genomic query region).To minimize chances that any genetic variants will be missed, it is a good idea to overlap amplicons by at least 25 bp. Figure 3 depicts PCR primer and amplicon design.
Follow PCR Primer Design Rules and Test Primers
PCR primer design for HRM-based screening follows the same general guidelines that apply to other PCR applications (See General PCR Primer Design Guidelines). Applied Biosystems® Primer Express® Software is an ideal tool for facilitating the primer design process. It is good practice to design three sets of primers for each amplicon and test them empirically to identify the most robust pair. Test for specific amplification of the target with minimal primer-dimer and nonspecific amplification using both traditional low-resolution melt curve analysis and agarose gel electrophoresis.
Step 2: Mutation Scanning Using HRM Analysis
Mutation scanning by HRM analysis is a fairly simple, straightforward process. Here we briefly outline the steps of the procedure; step-by-step instructions can be found in the Applied Biosystems High Resolution Melting Getting Started Guide (P/N 4393102).
DNA Sample Quality and Quantity
Because HRM is such a sensitive technique, try to limit experimental variability by using similar mass amounts of DNA in a comparable volume of low-salt buffer for each PCR reaction. With Applied Biosystems® MeltDoctor™ reagents, 20 ng DNA/reaction is recommended. Salts and other chemicals carried over from DNA isolation can also affect melt profiles; thus, it is important to use a DNA isolation technique with minimal contaminant carryover, and to use the same technique to obtain DNA from all samples and controls.
Replicates and Controls
As with most PCR-based techniques, it is good practice to run at least three replicates of each amplification reaction along with no-template controls. Likewise, if possible, include a known-variant control for each genomic region. Since most samples are likely to be wild type, the inclusion of one genomic variant provides evidence that mutations are detectable in each of the genomic regions queried. To enhance the reliability of samples identified as genetic variants, Applied Biosystems® High Resolution Melt Software v2.0 for the 7900HT System, High Resolution Melt Software v3.0 for the StepOne™, StepOnePlus™, and 7500 Fast Systems, and ViiA™ 7 Software HRM module for the ViiA™ 7 System can be directed to use data from up to 5 different wild type reactions for sample comparison.
PCR Amplification
HRM analysis is complicated by nonspecific amplification; thus, in addition to choosing optimal PCR primer pairs for each amplicon, PCR reagents that are specifically designed for HRM analysis can offer advantages in terms of accuracy, specificity, and reproducibility. MeltDoctor™ HRM reagents were developed for high-resolution melt analysis. They employ MeltDoctor™ HRM dye, a stabilized form of the SYTO® 9 dye—a next-generation dsDNA-binding dye developed by Molecular Probes that delivers sharp, clean melt profiles. Simplest to use is MeltDoctor™ HRM Master Mix; it requires only the addition of template DNA and a PCR primer pair before starting the PCR. When more flexibility in reactions is needed, the MeltDoctor™ HRM Reagent Kit provides enzyme, fluorescent dye, and dNTP mix separately. Both reagent sets employ hot-start–enabled DNA polymerase, minimizing nonspecific product formation and enabling reactions to be set up at room temperature. Plan experiments to minimize pipetting by combining PCR primers and other reagents into master mixes whenever possible. We recommend using 200 nM of each PCR primer and 20 ng of genomic DNA in 20 μL PCR reactions.
Amplify reactions for 40 cycles to produce enough material for melt analysis. Ideally, the amplification should be run on a real-time instrument to facilitate troubleshooting if that becomes necessary; however, since no data are collected during amplification, the PCR cycling can be done in a simple thermal cycler if desired. The melt analysis (dissociation), however, must be performed in an HRM-capable real-time PCR instrument.
Table 1. HRM PCR Primer Design Guidelines.
| Target Sequence | <250 bp* |
| Primer Length | ~20 nt |
| Primer Melt Temperature (Tm) | 58–62˚C, optimal 60˚C |
| Forward and Reverse Primer Tm Difference | <2˚C |
| GC Content | 30–80% |
| GC Clamp | Maximum of 2 G or C nucleotides in the last 5 nt at 3’ end |
| Avoid Runs of Identical Nucleotides (Especially G) | |
| Avoid Complementarity Within or Between Primers | |
| Avoid Primers with Homology to Other Targets |
HRM Analysis
During the HRM stage of the melt curve, double-stranded amplicons slowly denature, releasing bound dsDNA-binding dye. The real-time PCR instrument measures this decrease in fluorescence signal, and the HRM software plots fluorescence signal over temperature. Figure 4 shows an example of HRM data generated from amplification of 4 replicates of 24 samples using PCR primers for a 121 bp region of the NAT2 gene. Figure 4A shows the raw melt curves. Next, Applied Biosystems® HRM Software v2.0 scales the data by aligning the fluorescence values for 100% and 0% duplexed amplicons. Users can check and adjust the temperature cutoffs associated with these fluorescence values if necessary. The aligned melt curve graph is shown in Figure 4B. The melt (dissociation) curves from the variant DNA (red lines) can be distinguished from the many wild type samples in this view, but they become even more apparent when the software generates a difference plot by displaying the difference in fluorescence between a reference sample and the other melt curves, as shown in Figure 4C.
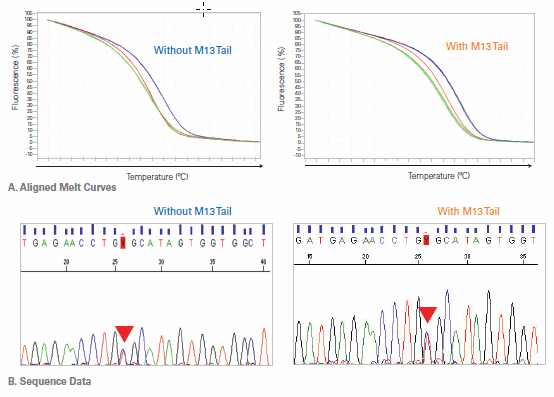
Figure 2. M13 Tailing of PCR Primers Changes the Melt Profile, but Not the Result. (A) HRM analysis data are shown for the same target amplified using PCR primers with and without the M13 universal DNA sequencing tail appended to the 5’ ends. In both sets of data, the melt profiles of homozygous wild type, homozygous mutant, and heterozygous can be distinguished from each other, but since the amplicons have different sequences, the curves are slightly different. (B) Sanger sequencing data from the heterozygote PCR product shown in Panel A.
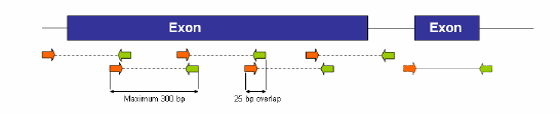
Figure 3. PCR Primer and Amplicon Design for Mutation Scanning via HRM. Design primers to query up to 250 bp of genomic target sequence per amplicon. Include at least 25 bp of overlap between amplicons to reduce the chance of missing genetic variants between amplicons.
Step 3: Sequencing the Genetic Variants Identified
PCR products found to contain mutation(s) by HRM scanning can be further analyzed via Sanger sequencing. In many cases, PCR products from HRM analyses need only be diluted prior to sequencing; typically no purification is needed. This is because, for amplicons smaller than ~300 bp, the DNA must be diluted >1:20 to obtain an appropriate template concentration for the sequencing reaction. This dilution reduces the concentration of leftover PCR primers and dNTPs to negligible levels in terms of the sequencing reaction.
For sequencing HRM reaction products, we recommend fast resequencing using the BigDye® Terminator v1.1 Cycle Sequencing Kit, which is well suited for short PCR products with rapid electrophoresis run modules (see Figure 5). In addition, the v1.1 kit was designed for optimal base calling adjacent to the sequencing primer, to avoid missing the first few bases of sequence.
Fast, Economical Mutation Scanning Followed by Rapid Sequencing
Compared to previous-generation mutation scanning methods such as dHPLC, DGGE, and direct DNA sequencing, HRM offers a significantly easier and less expensive method for quickly finding new genetic variations. Applied Biosystems now offers a complete set of reagents and software designed specifically for HRM applications such as mutation scanning, methylation analysis, and genotyping. The software and reagents were designed for use on Applied Biosystems® StepOne™, StepOnePlus™, 7500 Fast, 7900HT Fast, and ViiA™ 7 Real-Time PCR Systems, providing an intuitive user interface and reliable instrument performance for complete confidence in your results. HRM is a nondestructive process, so HRM reaction products that are identified for further study can be sequenced directly—no additional amplification, and in many cases, no additional purification, is needed prior to sequencing. Furthermore, with Applied Biosystems® fast sequencing reagents and capillary electrophoresis instruments, including the new 3500 Series Genetic Analyzers, DNA sequencing can be completed in a single day. From optimized plastic consumables to enzymatic reaction kits, intuitive yet powerful software, and innovative instrumentation, Applied Biosystems has the tools you need for sensitive, rapid mutation scanning followed by Sanger DNA sequencing to confirm and characterize new genetic variants.
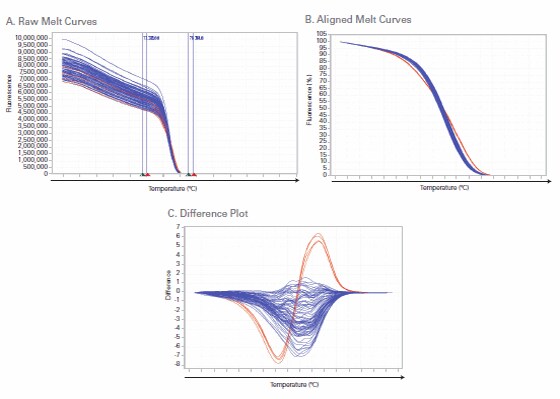
Figure 4. Example HRM Data. A 121 bp region of human NAT2 (N-acetyltransferase 2) gene was amplified from 4 replicates of 24 samples on the Applied Biosystems® 7500 Fast Real-Time PCR System. All images were generated with Applied Biosystems® HRM Software v2.0: (A) Raw melt (dissociation) curves; (B) Aligned melt curves; (C) Difference plot.

For Research Use Only. Not for use in diagnostic procedures.