Search
Methods for Detecting Protein–RNA Interactions
RNA electrophoretic mobility shift assay (EMSA)
The RNA electrophoretic mobility shift assay (RNA EMSA) is an in vitro technique used to detect protein–RNA interactions through changes in migration speed during gel electrophoresis. First, a labeled RNA probe is incubated with a protein sample (typically from a cell lysate) to initiate binding and formation of the interaction complex. The binding reaction is then separated via non-denaturing polyacrylamide gel electrophoresis. Like protein–DNA complexes, a protein–RNA complex migrates more slowly than a free RNA probe through a gel matrix. This causes a migration shift relative to the unbound RNA probe. Specificity is determined through a competition reaction, where excess unlabeled RNA is incubated in the binding reaction, resulting in a decrease in the shifted signal if the labeled and unlabeled RNA sequences compete for binding of the same protein. Alternatively, the protein–RNA complex may be crosslinked and the reaction run on a denaturing gel. Specificity is determined through visualization of a single shifted band. Traditionally, RNA probes are radioactively labeled for detection, although fluorescent and chemiluminescent detection is also possible. Non-radioactive RNA end-labeling techniques are limited, but more versatile biotin and fluorescent labeling methods are now available. For example, The Thermo Scientific LightShift Chemiluminescent RNA EMSA Kit provides a non-radioactive solution for studying RNA–protein interactions using EMSA.
| Strengths | Limitations |
|---|---|
|
|
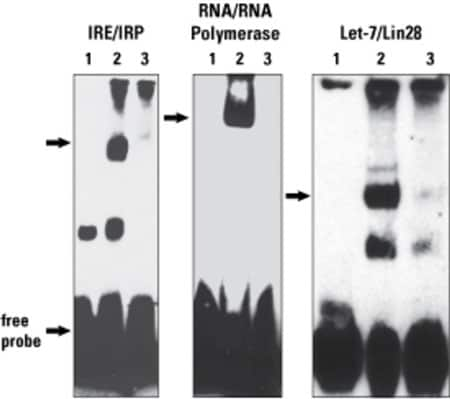
Chemiluminescent RNA EMSA using RNA probes biotinylated with the Pierce RNA 3' End Biotinylation Kit. Biotinylated RNA probes retain function and interact with RNA-binding proteins. Iron response element (IRE), RNA polymerase and Let-7 RNA probes were biotinylated using the Thermo Scientific Pierce RNA 3' End Biotinylation Kit. Each biotinylated RNA probe (5–10 nM) was tested for functionality in a RNA-binding reaction with 4–5 µg of cell extract using the LightShift Chemiluminescent RNA EMSA Kit to detect interactions with iron responsive protein (IRP), RNA polymerase or Lin28. Lane 1: free probe; Lane 2: binding reaction; Lane 3: binding reaction plus 100-fold excess of unlabeled probe. The presence of shifted bands (arrows) in lane 2 of each experiment shows RNA–protein complexes in the binding reactions, which are competed away by the addition of excess unlabeled RNA probe (lane 3).
Our 72-page Protein Interaction Technical Handbook provides protocols and technical and product information to help maximize results for protein interaction studies. The handbook provides background, helpful hints and troubleshooting advice for immunoprecipitation and co-immunoprecipitation assays, pull-down assays, far-western blotting and crosslinking. The handbook also features an expanded section on methods to study protein–nucleic acid interactions, including ChIP, EMSA, and RNA EMSA. The handbook is an essential resource for any laboratory studying protein interactions.
Contents include: Introduction to protein interactions, Co-immunoprecipitation assays, Pull-down assays, Far-western blotting, Protein interaction mapping, Yeast two-hybrid reporter assays, Electrophoretic mobility shift assays [EMSA], Chromatin immunoprecipitation assays (ChIP), Protein–nucleic acid conjugates, and more.

RNA pull-down assays selectively extract a protein–RNA complex from a sample. Typically, the RNA pull-down assay takes advantage of high affinity tags, such as biotin or azido-phosphine chemistry. RNA probes can be biotinylated, complexed with a protein from a cell lysate, and then purified using agarose or magnetic beads. Alternatively, the protein may be labeled, or the RNA–protein complex may be isolated using an antibody against the protein of interest. The RNA is then detected by northern blot or through RT-PCR analysis and the proteins detected by western blotting or mass spectrometry. We offer the Thermo Scientific Pierce Magnetic RNA-Protein Pull-Down Kit, which provides reagents to efficiently enrich RNA-binding proteins (RBPs) using RNA end-labeled with desthiobiotin and streptavidin magnetic beads.
| Strengths | Limitations |
|
|
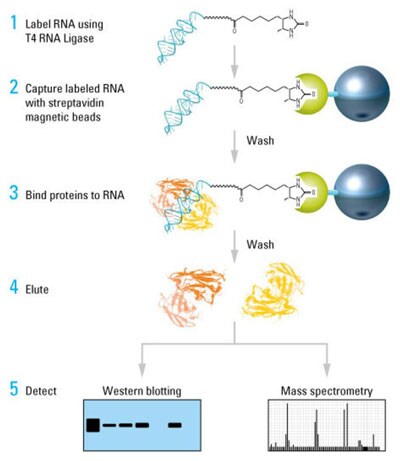
RNA–protein pull-down protocol. Diagram of the procedure for end-labeling RNA with desthiobiotin and then capturing and enriching specific RNA-binding proteins using the Pierce Magnetic RNA-Protein Pull-Down Kit.
The RNase protection assay (RPA) is a powerful method for detecting RNA and RNA fragments in cell extracts. Unlike northern blotting or RT-PCR analysis, RPA assays allow greater flexibility in the integrity of target RNA, requiring very short segments for hybridization and detection. RPA assays can also be used to map protein–RNA interactions. In this adaptation of the RPA, RNase H is used to cleave a target RNA molecule at a specific site hybridized with a DNA probe. If a protein is bound to the RNA at the target sequence, it will block probe hybridization, prevent cleavage by RNase H, and indicate a site of interaction between protein and RNA. RNase H requires only a four base pair hybrid with a DNA probe in order to cleave the RNA molecule of interest. Using many small probes allows the entire sequence of RNA to be mapped for sites of interaction.
| Strengths | Limitations |
|
|
Fluorescent in situ hybridization (FISH/ISH) co-localization techniques require detection of both a RNA transcript and a protein of interest using RNA probes and antibodies. FISH/ISH detects the position and abundance of a RNA and protein in a cell or tissue sample. The readout is visual (usually imaged via microscopy), and a co-localized signal for both the RNA and protein of interest indicate possible complex formation. A labeled RNA probe must be generated for detection of a particular sequence of RNA, and the protein may be detected using antibody staining or fluorescent protein constructs.
| Strengths | Limitations |
|
|
Traditional FISH techniques using large oligonucleotide sequences labeled with one to five fluorophores are generally limited by high background and low sensitivity due to nonspecific binding and insufficient signal amplification. Invitrogen ViewRNA assays incorporate a proprietary probe set design and branched DNA (bDNA) signal amplification technology. A target-specific probe set of approximately 20 oligonucleotide pairs hybridizes to the target RNA of interest. Signal amplification is achieved through specific hybridization of adjacent oligonucleotide pairs to bDNA structures, which are formed by pre-amplifiers, amplifiers, and fluorochrome-conjugated label probes. This methodology results in greater specificity, lower background and higher signal-to-noise ratios.
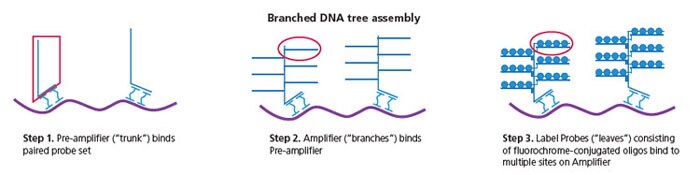
Formation of bDNA. A schematic depicting how signal amplification is achieved with bDNA technology.
Select products
Recommended reading
- Mendes ND et al. (2009) Current tools for the identification of miRNA genes and their targets. Nucleic Acids Research 37:2419–2433.
- Gunzl A, Bindereif A (1999) Oligonucleotide-targeted RNase H protection analysis of RNA-protein complexes. Methods Mol Biol 118:93–103.
For Research Use Only. Not for use in diagnostic procedures.