Search
Real-time PCR has become the method of choice for quantitating gene expression.
It is simple and robust, and offers benefits such as a 9-log dynamic range, rapid results, sensitivity, specificity, and reproducibility that make other methods obsolete. Traditionally, the first steps in real-time PCR are to isolate and quantitate RNA. However, these steps can require significant time and hands-on manipulation, and risk sample loss, especially from small samples. The Ambion® Cells-to-CT™ Kits provide a simple and rapid method for preparing cell lysates that can be used directly in gene expression analysis (see Figure 2). Using the Cells-to-CT methodology, relative gene expression is measured by real-time PCR directly from cell lysates, eliminating the need to purify and quantitate RNA prior to amplification. Here, we demonstrate that relative gene expression data obtained from real-time PCR using the Cells-to-CT workflow is equivalent to that obtained using purified RNA quantitated by spectrophotometry prior to real-time PCR, but with significantly less time and effort required and greater sensitivity.
Real-Time PCR for Target Quantitation
Absolute Quantitation
Initial real-time PCR quantitation strategies relied heavily on accurate mass amounts of input template. The so-called “absolute quantitation” method involves creating a standard curve of amplification of known amounts of target, and then using the observed CT values from different known input amounts for comparison to samples containing unknown target amounts. Other researchers have simply used the same mass amount of each sample and compare the “raw” CT values from real-time PCR for the desired target; this is referred to as “mass normalization”.
Relative Quantitation
Compared to absolute quantitation, relative quantitation is significantly quicker and easier, and is thus becoming the preferred method for real-time PCR data interpretation. Relative quantitation of gene expression involves reverse transcription (RT) followed by real-time PCR of a control target (with constant, characterized expression levels) to normalize expression of unknown targets; these control targets are often called “normalizers”. (see Selecting Endogenous Controls for Real-time PCR.) With the right real-time PCR reagents and instruments, both the target-of-interest and the endogenous control can be amplified in a single “duplex” reaction, adding to the simplicity of this method.
Simplify Relative Quantitation Using Ambion® Cells-to-CT™ Methodology
Ambion Cells-to-CT technology enables real-time PCR of cultured cell samples without isolating RNA. Potential problems such as sample mix-up and RNA degradation are minimized, and samples are not subject to RNA quantitation errors. With Cells-to-CT kits, cell lysates are ready for reverse transcription (RT) after a two-step, 7-minute lysis procedure at room temperature that includes an option for DNase treatment to simultaneously remove genomic DNA. Samples can contain 10–100,000 cells, and prepared lysates can be used in RT reactions without quantitation of any kind. Customers sometimes ask how they can quantitate the RNA in their Cells-to-CT lysates—the answer is that it is not necessary and here we present data to support this claim (see also "Skipping RNA Quantitation: Common Concerns", in sidebar).
RNA Concentrations from Small Samples Cannot be Measured Using Spectrophotometry or RiboGreen Reagent
Comparable Gene Expression Results from Known RNA Mass Amounts or Cells-to-CT Lysates
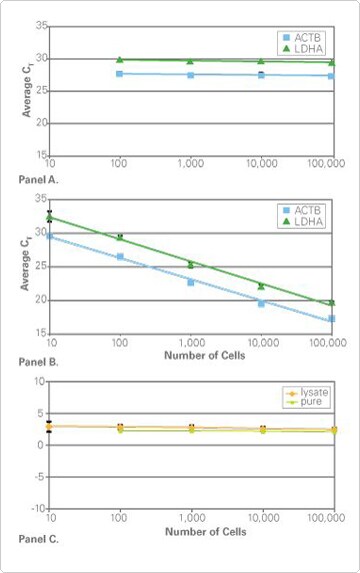
Figure 1. Equivalent Gene Expression Results With or Without Quantitating RNA Prior to Real-Time PCR. Panel A. Real-Time PCR Results from RNA Mass Normalization. Using the RNA concentrations measured with RiboGreen Reagent, equal mass amounts (383 pg) of RNA isolated from the indicated number of cells was reverse transcribed and the cDNA was used in duplicate real-time PCRs using TaqMan® Gene Expression Assays for ACTB (assay ID#HS03023889_g1) and LDHA (assay ID#HS00855332_g1) and TaqMan Gene Expression Master Mix. The observed CTs are equivalent, demonstrating that equal mass amounts of RNA in RT-PCR produce equivalent real-time PCR results. Panel B. Real-Time PCR Results from TaqMan Cells-to-CT™ Lysates. The indicated numbers of cells were processed using the TaqMan Gene Expression Cells-to-CT Kit and lysates were used directly in duplicate real-time PCRs. The results demonstrate that lysis, RT and real-time PCR reactions are linear over 10–100,000 input cells, and that adding a specific amount of RNA into the RT reaction is unnecessary. Panel C. Equivalent ΔCT Results. The indicated numbers of cells were processed either using the TaqMan Gene Expression Cells-to-CT Kit (orange line) or for RNA purification (green line). ΔCT values were calculated by subtracting the ACTB CT values from the LDHA CT values.
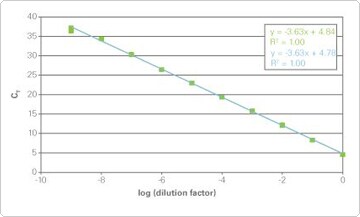
Figure 2 (sidebar). 9-log Dynamic Range of Real-Time PCR. Reverse transcription reactions from mouse NIH/3T3 cell lysate (green) or purified RNA (blue) were used for real-time PCR. 9-logs of template were efficiently detected in the presence of Cells-to-CT lysate.
An endogenous control, also commonly called a normalizer, reference gene, or housekeeping gene, is a gene whose expression can serve as a benchmark for comparison to that from the gene-of-interest. It should be expressed at the same level in all of the samples to be compared. Commonly used normalizer sequences include PPIA, TNP, GAPDH, and 18S. (For a complete list, refer to Applied Biosystems TaqMan® Express Human Endogenous Control Plate). To select the best reference genes for your system:
1. Identify candidate normalizer genes (for example, the TaqMan Express Human Endogenous Control Plate).
2. Obtain a representative group of samples for your experiments (for example, treated and untreated cell cultures).
3. Perform real-time RT-PCR with assays for all of the candidate normalizer genes with equal amounts of all the samples.
4. The best normalizer is one that:
a) is sufficiently abundant in every sample for reliable quantitation.
b) shows tight covariance among a panel of candidate endogenous controls, between your samples. Statistical tools such as StatMiner™ Software and geNorm are available for this analysis
Skipping RNA Quantitation: Common Concerns
Some people find it hard to get used to the idea that, with Cells-to-CT technology, quantitating RNA before RT-PCR is unnecessary. Here, Richard Fekete and David Keys, Research and Development scientists at Applied Biosystems, address some of these concerns.
- One issue is simply that researchers are accustomed to quantitating RNA prior to real-time PCR. After a few experiments with Cells-to-CT that eliminate the time and effort of isolating and quantitating RNA, most people no longer have any reservations about leaving these steps behind. Like mass normalization, relative quantitation normalizes real-time PCR results, however, it is performed bioinformatically after the reaction rather than by adjusting RNA concentrations before the reaction. Additionally, since the ΔCT measurements are calculated relative to an internal biological control, they are less subject to measurement errors in RNA quantitation or cell counts.
- Researchers might be concerned that their experimental treatment either kills most of the cells or results in very slow growth; this might not be apparent from a ΔCT measurement that represents the difference in CT values from the experimental and normalizer (endogenous control) PCRs. This concern can be addressed fairly easily by simply comparing the CT values for the normalizer PCR across samples. Those reactions containing significantly fewer cells than the majority of the samples will exhibit a proportionally higher CT value for the normalizer PCR.
- Large experiments. Cells-to-CT lysates can be qualified for further studies by performing a pilot PCR for an endogenous control target.
- Staying within the dynamic range of both the reverse transcription and the real-time PCR may be a concern. However, all Cells-to-CT kits demonstrate lysis efficiency over a 4–log dynamic range of cell input from 10–100,0000 cells, and real-time PCR is well established to support over a 9–log dynamic range (Figure 2, below left). Therefore, by keeping cell input at 100,000 cells/sample or less, the Cells-to-CT strategy is equally appropriate for evaluating highly expressed genes from as many as 100,000 cells or samples that are too small for traditional RNA purification and quantitation methodologies.