Search
Proteintech
PSMC2 Polyclonal Antibody
{{$productOrderCtrl.translations['antibody.pdp.commerceCard.promotion.promotions']}}
{{$productOrderCtrl.translations['antibody.pdp.commerceCard.promotion.viewpromo']}}
{{$productOrderCtrl.translations['antibody.pdp.commerceCard.promotion.promocode']}}: {{promo.promoCode}} {{promo.promoTitle}} {{promo.promoDescription}}. {{$productOrderCtrl.translations['antibody.pdp.commerceCard.promotion.learnmore']}}
Product Details
14905-1-AP
Species Reactivity
Published species
Host/Isotype
Class
Type
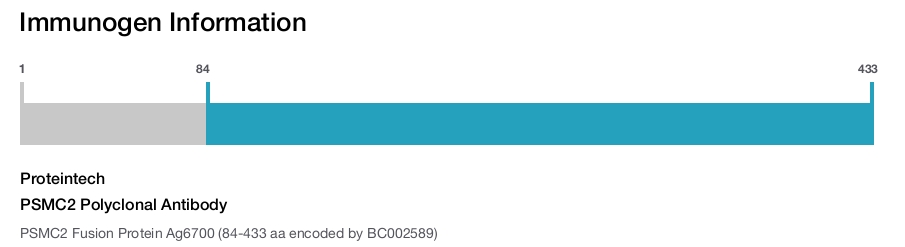
Immunogen
Conjugate
Form
Concentration
Amount
Purification
Storage buffer
Contains
Storage conditions
Shipping conditions
Product Specific Information
Immunogen sequence: KQTLQSEQP LQVARCTKII NADSEDPKYI INVKQFAKFV VDLSDQVAPT DIEEGMRVGV DRNKYQIHIP LPPKIDPTVT MMQVEEKPDV TYSDVGGCKE QIEKLREVVE TPLLHPERFV NLGIEPPKGV LLFGPPGTGK TLCARAVANR TDACFIRVIG SELVQKYVGE GARMVRELFE MARTKKACLI FFDEIDAIGG ARFDDGAGGD NEVQRTMLEL INQLDGFDPR GNIKVLMATN RPDTLDPALM RPGRLDRKIE FSLPDLEGRT HIFKIHARSM SVERDIRFEL LARLCPNSTG AEIRSVCTEA GMFAIRARRK IATEKDFLEA VNKVIKSYAK FSATPRYMTY N (84-433 aa encoded by BC002589)
Target Information
Proteolytic degradation is critical to the maintenance of appropriate levels of short-lived and regulatory proteins as important and diverse as those involved in cellular metabolism, heat shock and stress response, antigen presentation, modulation of cell surface receptors and ion channels, cell cycle regulation, transcription, and signalling factors. The ubiquitin-proteasome pathway deconstructs most proteins in the eukaryotic cell cytosol and nucleus. Others are degraded via the vacuolar pathway which includes endosomes, lysosomes, and the endoplasmic reticulum. The 26S proteasome is an ATP-dependent, multisubunit (~31), barrel-shaped molecular machine with an apparent molecular weight of ~2.5 MDa. It consists of a 20S proteolytic core complex which is crowned at one or both ends by 19S regulatory subunit complexes. The 19S regulatory subunits recognize ubiquitinated proteins and play an essential role in unfolding and translocating targets into the lumen of the 20S subunit. An enzymatic cascade is responsible for the attachment of multiple ubiquitin molecules to lysine residues of proteins targeted for degradation. Several genetic diseases are associated with defects in the ubiquitin-proteasome pathway. Some examples of affected proteins include those linked to cystic fibrosis, Angelman's syndrome, and Liddle syndrome.
For Research Use Only. Not for use in diagnostic procedures. Not for resale without express authorization.
Bioinformatics
Protein Aliases: 26S protease regulatory subunit 7; 26S proteasome AAA-ATPase subunit RPT1; 26S proteasome regulatory subunit 7; mammalian suppressor of sgv-1 of yeast; MGC3004; protease 26S subunit 7; proteasome (prosome, macropain) 26S subunit, ATPase, 2; Proteasome 26S subunit ATPase 2; putative protein product of Nbla10058; testis secretory sperm-binding protein Li 197a; unnamed protein product
Gene Aliases: MSS1; Nbla10058; PSMC2; RPT1; S7
UniProt ID: (Human) P35998, (Rat) Q63347, (Mouse) P46471
Entrez Gene ID: (Human) 5701, (Rat) 25581, (Mouse) 19181

Performance Guarantee
If an Invitrogen™ antibody doesn't perform as described on our website or datasheet,we'll replace the product at no cost to you, or provide you with a credit for a future purchase.*
Learn more
We're here to help
Get expert recommendations for common problems or connect directly with an on staff expert for technical assistance related to applications, equipment and general product use.
Contact tech support