If you work in a clinical research la b, you might be familiar with the manual process of comparing your data from your NGS platform with data from your Sanger Sequencing system. It is a time consuming process that probably requires you to rearrange your calendar depending on the quality of your data. Well, good news! NGS confirmation is a snap with one of the newest software solutions from Applied Biosystems®.
b, you might be familiar with the manual process of comparing your data from your NGS platform with data from your Sanger Sequencing system. It is a time consuming process that probably requires you to rearrange your calendar depending on the quality of your data. Well, good news! NGS confirmation is a snap with one of the newest software solutions from Applied Biosystems®.
At ASHG 2015 in Baltimore, Venkatesh Moktali led two live demo sessions at the Thermo Fisher Scientific booth, giving customers a quick glimpse of how these new software solutions can streamline their NGS confirmation workflow. We managed to catch up with Venkatesh for a few minutes before his demo to talk about what he is bringing to the table.
The new workflow using the Next Generation Sequencing Confirmation (NGC) module consists of five simple steps:
- Import your sanger “.ab1” files from any Capillary electrophoresis instrument
- Create a Reference to map the variants and upload your vcf file containing NGS variants
- Organize your data into Amplicons for grouping of reads
- Set parameters for Trimming, Filtering and Alignment.
- Review your settings and run the analyses.
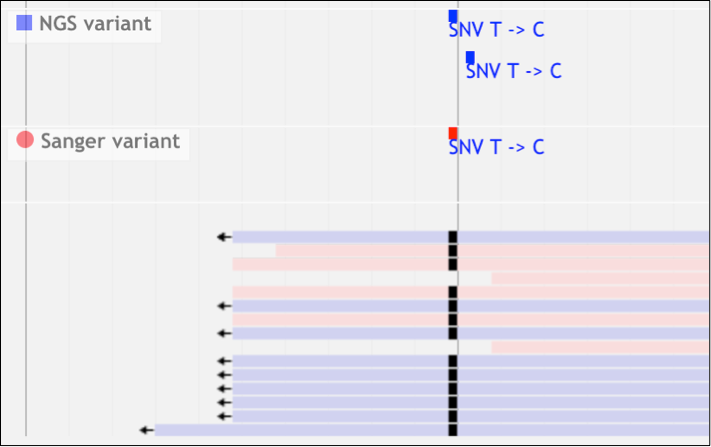
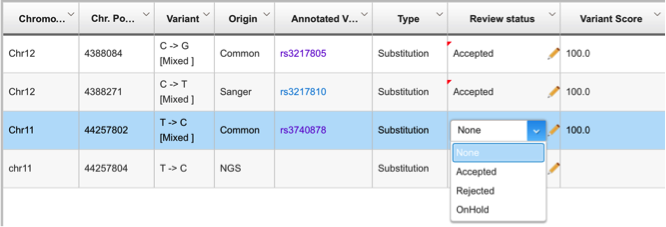
The NGC module shows the comparison of variants from each of the sequencing platforms through an overview pane that puts the variants into two tracks supported by Sanger reads piled up in another track. The module also lets you review your variants and choose the highly confident and confirmed variants through a review table:
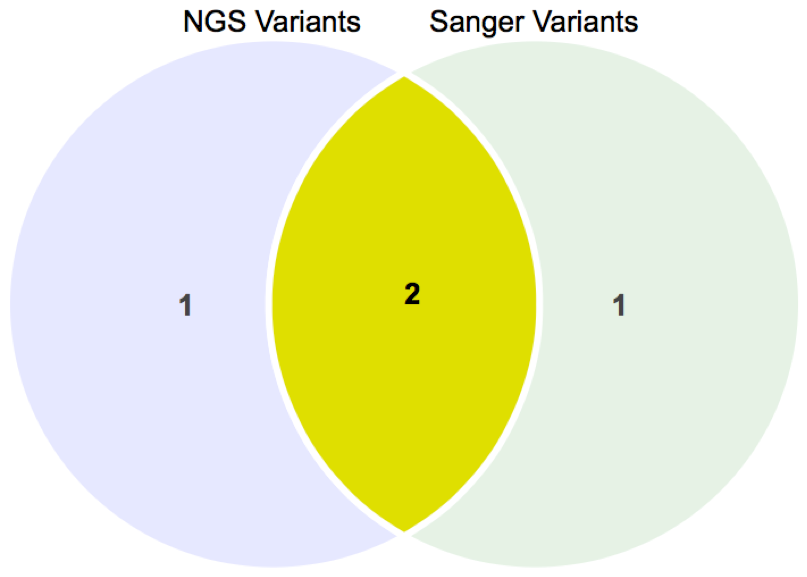
Finally, you can view your confirmed variants and total variants in a beautiful Venn diagram that gives you a snapshot of your results. The results can also be downloaded in vcf format that consists of all your review results.
 Looks easy enough? Give it a try! The NGC module is now FREE on Thermo Fisher Cloud.
Looks easy enough? Give it a try! The NGC module is now FREE on Thermo Fisher Cloud.




Leave a Reply