 Y-STR analysis has been successfully used in a variety of criminal cases, such as sexual assault cases, missing person identification, familial searching, and generating investigative lead for paternal biogeographic ancestry information. Because of Y-STR loci’s relatively high mutation rates (~10-4to ~10-2), it is not uncommon to observe that two related Y-STR profiles (e.g., father and son) have different genotypes at one or more loci. Therefore, forensic scientists are in need of a method to determine if two Y-STR profiles are from the same male lineage or not.
Y-STR analysis has been successfully used in a variety of criminal cases, such as sexual assault cases, missing person identification, familial searching, and generating investigative lead for paternal biogeographic ancestry information. Because of Y-STR loci’s relatively high mutation rates (~10-4to ~10-2), it is not uncommon to observe that two related Y-STR profiles (e.g., father and son) have different genotypes at one or more loci. Therefore, forensic scientists are in need of a method to determine if two Y-STR profiles are from the same male lineage or not.
There is a long answer, as well as a short answer. Ideally, just as in autosomal profile analysis, Y-STR profiles should be interpreted under a likelihood ratio framework, which generates prosecution and defense hypotheses with fixed pedigree relationships for the evidence profile, and calculates the pedigree likelihood ratio given hypotheses according to the family structure and mutation rates. The details of this method have been described in Ge et al. [1]. Unfortunately, this method requires pedigree information for each hypothesis, which may not be practically available in many cases. Besides, the algorithm is too cumbersome for most analysts to implement.
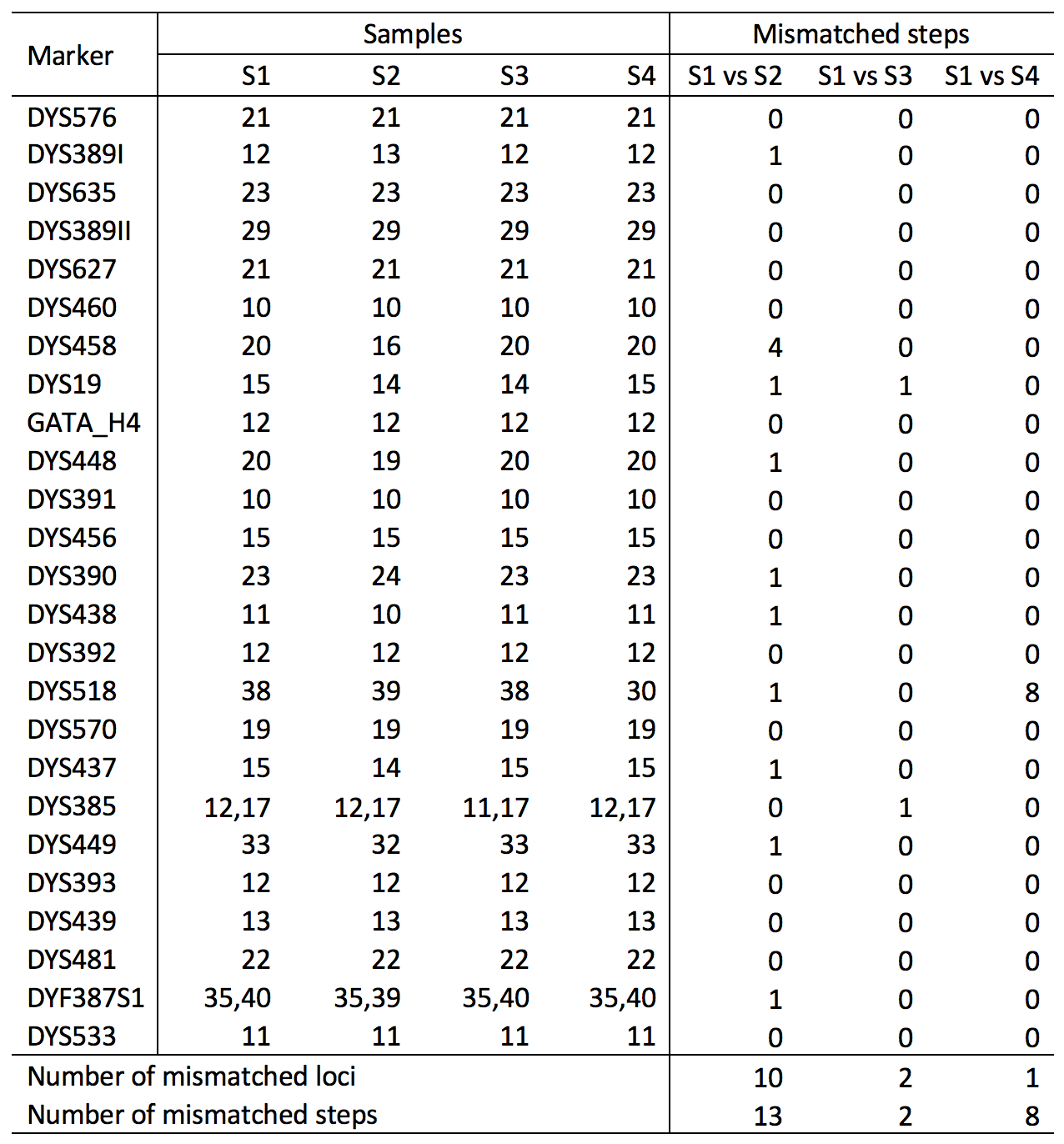
A more convenient approach is to count how many mismatched steps exist between two profiles. Based on the study by Liu et al. [2], assuming no genotyping or human error, two full YfilerTMPlus profiles with ≤5 mismatched steps are more likely from the same lineage, in which “step” is defined as the repeat different between two alleles (e.g., 10 and 12 have 2 mismatched steps). The following example demonstrates how this approach works. In this example, S1 and S3 have only 2 mismatched steps, which means they are more likely from the same male lineage (i.e., 2≤5). S1 and S2 are more likely not related (i.e., 13>5). S1 and S4 are determined as not related based on this approach. However, in this scenario, it would be advisable to double check if the alleles at the mismatch locus (DYS518) was mistyped, as it appears a genotyping error may have occurred at DYS518 and the genotyping error rate is generally at the level of 0.1%.
In general, the above approach gives a quick way to answer if two Yfiler Plus haplotypes are from the same male lineage. The threshold may be different for different commercial kits. There is no guarantee that this approach would always give you the right decision. Fortunately, two unrelated profiles, in most cases (i.e., >99.975%), will have more than 5 mismatched steps; and, for almost all close relatives, the mismatched steps are ≤5 (i.e., 99.999997% father-son, 99.999984% grandfather-grandson, 99.999950% uncle-nephew, and 99.999878% cousin have mismatched steps ≤5). This is an approach you might use to make a quick decision on whether two male profiles share the same lineage with high confidence.
- Ge J, Eisenberg A, Yan J, Chakraborty R, Budowle B. Pedigree likelihood ratio for lineage markers. Int J Legal Med. 2011 Jul;125(4):519-25.
- Liu H, Li X, Mulero J, Carbonaro A, Short M, Ge J. A convenient guideline to determine if two Y‐STR profiles are from the same lineage. Electrophoresis. 2016 Jul 1;37(12):1659-68.
For Research, Forensic, or Paternity Use Only. Not for use in diagnostic procedures.
Leave a Reply