Search
Citations et références (1)
Invitrogen™
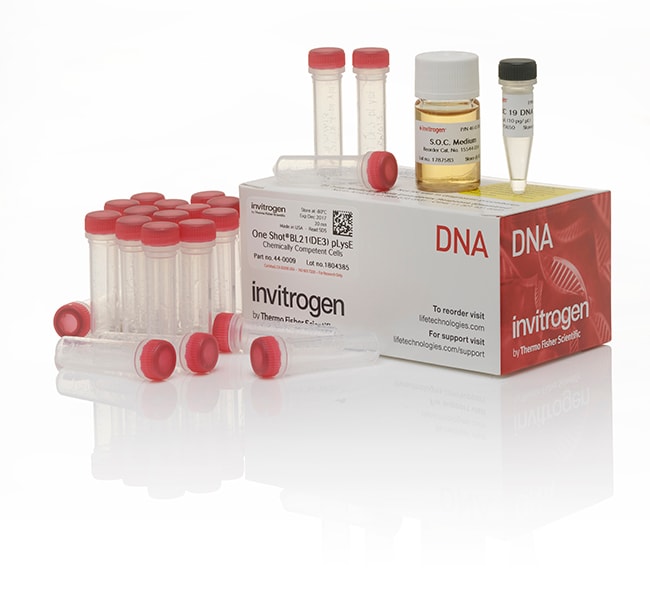
One Shot™ BL21(DE3)pLysE Chemically Competent E. coli
Les cellules E. coli BL21(DE3)pLysE sont idéales pour une utilisation avec des vecteurs d’expression basés sur le promoteur T7. ToutAfficher plus
| Référence | Quantité |
|---|---|
| C656503 | 21 x 50 μl |
Référence C656503
Prix (EUR)
428,65
Online Exclusive
471,00Économisez 42,35 (9%)
Each
Quantité:
21 x 50 μl
Prix (EUR)
428,65
Online Exclusive
471,00Économisez 42,35 (9%)
Each
Les cellules E. coli BL21(DE3)pLysE sont idéales pour une utilisation avec des vecteurs d’expression basés sur le promoteur T7. Tout comme les cellules E. coli BL21(DE3)pLysS, les cellules BL21(DE3)pLysE transportent le lambda lysogène DE3. De plus, les cellules BL21(DE3)pLysE contiennent le plasmide pLysE qui exprime constitutivement le lysozyme T7. Le lysozyme T7 réduit l’expression basale des gènes cibles en inhibant l’ARN polymérase T7. Le plasmide pLysE dans les cellules BL21(DE3)pLysE exprime des niveaux plus élevés de lysozyme T7 que le plasmide pLyss dans les cellules BL21(DE3)pLysS.
La souche BL21(DE3)pLysE permet de contrôler de manière plus strict l’ARN polymérase T7 nécessaire lorsque la protéine recombinante qui doit être exprimée est toxique.
La souche BL21(DE3)pLysE permet de contrôler de manière plus strict l’ARN polymérase T7 nécessaire lorsque la protéine recombinante qui doit être exprimée est toxique.
Usage exclusivement réservé à la recherche. Ne pas utiliser pour des procédures de diagnostic.
Spécifications
Résistance aux antibiotiques des bactériesYes (Chloramphenicol)
Sélection bleue / blancheNon
Clonage d’ADN méthyléNon
Contient l’épisome FAbsence d’épisome F’
Compatibilité à haut débitNon compatible avec le haut débit (manuel)
Améliore la qualité des plasmidesNon
Improves Protein StabilityYes (lon, ompT)
Improves RNA StabilityYes (pLysE)
Préparation de l’ADN non méthyléYes (dcm)
Gamme de produitsOne Shot
Type de produitCellule compétente
Quantité21 x 50 μl
Réduit la recombinaisonNon
Conditions d’expéditionDry Ice
Résistant au phage T1 (tonA)Non
Toxic ProteinsNo
Niveau d’efficacité de la transformationEfficacité du sous-clonage (10^6-10^7 cfu⁄ µg)
FormatTube
AccélérateurT7
EspècesE. coli
Unit SizeEach
Contenu et stockage
• BL21 Star (DE3)pLysE Chemically Competent E. coli (21 x 50 μL); store at –80°C
• pUC19 DNA (1 x 50 μL); store at –20°C
• S.O.C. Medium (6 mL); store at 4°C or room temperature
• pUC19 DNA (1 x 50 μL); store at –20°C
• S.O.C. Medium (6 mL); store at 4°C or room temperature
Foire aux questions (FAQ)
My gene of interest is toxic to bacterial cells. Are there any precautions you can suggest?
I'm trying to express my protein using a bacterial expression system. How do I know if I'm seeing degradation of my protein or if what I’m seeing is codon usage bias?
I'm trying to express my protein using a bacterial expression system and am getting inclusion bodies. What should I do?
I'm getting low protein yield from my bacterial expression system. What can I do to improve this?
My cells are growing very slowly, and I'm not getting any protein expression from my baterial expression system. What can I do to fix this?
Citations et références (1)
Citations et références
Abstract
A novel group of oleosins is present inside the pollen of Arabidopsis.
Journal:J Biol Chem
PubMed ID:11929861
In plants, subcellular triacylglycerol granules in seeds (oil bodies) and floral tapetum (tapetosomes) are stabilized by amphipathic structural protein called oleosin. We hereby report a novel group of oleosins that is present inside the pollen of Arabidopsis thaliana. We have used the conserved sequence of oleosins to locate, via the