
1 in 10 people lack access to safe water worldwide, and about half the U.S. population lives without access to safe water. As the human population continues to grow, access to safe water is of utmost importance. Water is needed not only for human consumption but also for livestock and agricultural use. Regions experiencing water scarcity turn to using water treatment facility to reuse wastewater for agricultural or even human use. Water treatment is not a well-researched process, but recent studies show that many treatment processes allow reintroduction of antibiotics, antibiotic resistance genes and resistant organisms back into the environment3,4,5,6.
Commonly used to treat and prevent deadly infections, antibiotics are widely used all around the globe and are one of the most overprescribed drugs. Over-prescription of drugs and increasing applications in agriculture and livestock is pushing the evolution of microbes to be resistant to one or more antibiotic, eventually giving rise to multi-drug resistant (MDR) strains1. Contamination of the environment with antibiotics and resistance genes is becoming an increasing threat to global health3.
Evolution of microbes: How do microorganisms gain resistance to drugs?
Bacteria can gain resistance genes via horizontal gene transfer, which occurs between two bacteria or possibly via direct uptake of antibiotic resistance genes present in the environment2,3. Water environments are somewhat favorable for transfer for resistance genes3. Even though MDR organisms are found in hospitals, these organisms today can also be found in polluted water sources and the environment. Antibiotics and antibiotic resistance genes from these microbes can potentially bypass water treatment facilities and eventually make it back into the environment and back into our bodies.
Antibiotics have saved lives since their introduction in the 1940s, however, according to the CDC in the United States alone, antibiotic resistance is responsible for more than 2 million infections and 23,000 deaths at a direct cost of $20 billion1. Globally there is an overall decline in the effectiveness of antibiotics with a rise in resistance to first-line and last-resort antibiotics. Why is that so? One mi ght think – I don’t take antibiotics, so how might I be affected? Even if we may not directly take antibiotics, these drugs are widely used in agriculture and aquaculture, and we are exposed to antibiotics when we interact/come in contact or consume meats, eggs, and milk from these animals1. As the human population continues to grow, so does the demand for animal protein. Farmers not only use antibiotics to prevent diseases, but also to promote growth – a topic that is controversial1. Interestingly, wastewater treatment plants can be another unexpected source of contact with antibiotics and antibiotic resistance genes, as water-scarce countries and regions turn to the use of treated water for human consumption or agricultural use4.
ght think – I don’t take antibiotics, so how might I be affected? Even if we may not directly take antibiotics, these drugs are widely used in agriculture and aquaculture, and we are exposed to antibiotics when we interact/come in contact or consume meats, eggs, and milk from these animals1. As the human population continues to grow, so does the demand for animal protein. Farmers not only use antibiotics to prevent diseases, but also to promote growth – a topic that is controversial1. Interestingly, wastewater treatment plants can be another unexpected source of contact with antibiotics and antibiotic resistance genes, as water-scarce countries and regions turn to the use of treated water for human consumption or agricultural use4.
The First Line of Defense: How effective is a water treatment facility?
 Wastewater from households, hospitals, and pharmaceutical companies may carry antibiotics, deadly microbes, antibiotic resistance genes, and harsh chemicals and drugs3,6,7. In water-stressed regions, wastewater is treated at wastewater treatment plants and reused not only for human use but also for use in agricultural irrigation3,4,6,7. Since water environments are favorable for the transfer of resistance genes, it is important to understand how the presence of various antibiotics, drugs, and chemicals may affect microbial communities and the transfer of resistance genes. Wastewater from the manufacture of pharmaceutical products is a complex mixture of chemicals and microbes with high levels of organic matter, and anaerobic treatment is a favorable alternative3. Aydin et al. (2016) use quantitative PCR (qPCR) and next generation sequencing technology to study how complex microbial communities interact in an anaerobic bioreactor and the occurrence of antibiotic resistance genes3. They used qPCR to quantify antibiotic resistance genes and 16S rRNA sequencing on the Ion PGM™ sequencer to identify the diversity of the microbes. They observed that anaerobic treatment is unable to completely remove antibiotic resistance genes from wastewater, and the presence of multiple antibiotics lead to an increase in the occurrence of antibiotic resistance genes. They also suggest that quantitative real-time PCR may be a fast, sensitive and specific method for detection and quantification of resistant genes in water and sludge samples.
Wastewater from households, hospitals, and pharmaceutical companies may carry antibiotics, deadly microbes, antibiotic resistance genes, and harsh chemicals and drugs3,6,7. In water-stressed regions, wastewater is treated at wastewater treatment plants and reused not only for human use but also for use in agricultural irrigation3,4,6,7. Since water environments are favorable for the transfer of resistance genes, it is important to understand how the presence of various antibiotics, drugs, and chemicals may affect microbial communities and the transfer of resistance genes. Wastewater from the manufacture of pharmaceutical products is a complex mixture of chemicals and microbes with high levels of organic matter, and anaerobic treatment is a favorable alternative3. Aydin et al. (2016) use quantitative PCR (qPCR) and next generation sequencing technology to study how complex microbial communities interact in an anaerobic bioreactor and the occurrence of antibiotic resistance genes3. They used qPCR to quantify antibiotic resistance genes and 16S rRNA sequencing on the Ion PGM™ sequencer to identify the diversity of the microbes. They observed that anaerobic treatment is unable to completely remove antibiotic resistance genes from wastewater, and the presence of multiple antibiotics lead to an increase in the occurrence of antibiotic resistance genes. They also suggest that quantitative real-time PCR may be a fast, sensitive and specific method for detection and quantification of resistant genes in water and sludge samples.
While Aydin et al. studied anaerobic water treatment, Al-Jassim et al. studied conventional water treatment. Al-Jassim et al. (2015) studied the removal efficiency of microbial contaminants in a local wastewater treatment plant over the course of a year4. Al-Jassim et al. (2015) used the Applied Biosystems™ 7900HT Fast Real-Time PCR system to determine the number of gene copies present in their samples. They used the Ion PGM sequencer to determine microbial communities by assessing 16S rRNA from their samples. Similar to Aydin et al. (2016), this group too observed the presence of antibiotic resistance genes and antibiotic resistant bacteria in conventionally treated water.
One might think that minute quantities of antibiotics present in treated water ought to not be of concern. But what we need to understand is exactly how negligible of an amount can promote the transfer of resistance genes. Jutkina et al. (2016) developed an assay to determine minimal concentrations of antibiotics that drive transfer of resistance5. Their assay showed that very low concentrations of tetracycline, about 150 times below the minimum concentration that will inhibit the growth of the microbe promoted horizontal transfer of the multiple antibiotic resistance genes, with higher concentrations favoring more transfer and selection towards a tetracycline resistant phenotype5. Water treatment plants, processes, and removal efficiency is a poorly understood area. Research using more sensitive assays and instruments may help better understand and help identify ways to improve efficiencies to water treatment processes.
Back into the Environment and water sources: What happens to the water once it leaves the water treatment facility?

One third of gastrointestinal diseases and ~4% of all worldwide deaths occur due to contaminated drinking water and poor sanitation6. A variety of biological, physical and chemical indicators are used to assess efficiency of water treatment processes and most drinking water quality assessments do not directly identify various bacterial taxa present. Traditional water testing methods are primarily culture-based and less than 2% of bacterial species can be cultured in the lab8. Culture based methods take longer and miss unculturable bacteria that cannot be cultured in the lab. Although these traditional methods test for the presence or the number of bacterial cells. these methods provide limited information, if any, about the type of bacteria or strain present8.
Shaw et al. (2015) used the Ion PGM sequencer to perform 16S rRNA metagenomic amplicon sequencing to test water quality of two drinking water distribution systems6. This approach was able to detect various taxa present in water subjected to disinfection and post-disinfection before distribution. Not only were they able to detect various bacterial taxa, but they also accurately identified the presence of various bacteria communities. Water systems may be effective in reducing bacterial communities in treated disinfected water, but once the water leaves the plant and is distributed, bacterial communities can regrow. Hence, it is crucial to identify the taxa of the bacteria present in treated water before distribution. Their data also observed that microbial communities identified in samples with high amounts of disinfectant are more likely to be resistant species. Shaw et al. suggest that metagenomic amplicon sequencing can be a promising, rapid and inexpensive technique for identifying multiple taxa from complex biological samples6.
Let the microbial battlefield begin: If not antibiotics, how else can we combat these organisms?
The question now turns to – how we can combat MDR organisms or deadly bacterial diseases without the aid of antibiotics? Developing effective treatments is necessary to thoroughly understanding the organism, if and how it is evolving, and its level of drug-resistance.
Graham et al. (2016) used next-generation sequencing to sequence the entire genome of Neisseria gonorrhoeae from DNA extracted from the urine of 13 specimens already known to be positive for N. gonorrhoeae7. With the use of the Ion PGM sequencer, they were able to reconstruct the complete genome of N. gonorrhoeae from these samples. Sequencing the entire genome gave insights into novel antibiotic resistance markers that helped better understand the evolution and nature of N. gonorrhoeae, but may potentially be used for epidemiological surveillance and typing this organism7.
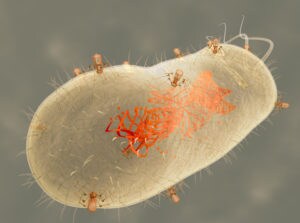 Shigella is a group of bacteria that cause Shigellosis, an infectious disease with a majority of Shigellosis cases occurring in developing countries. This bacterial pathogen is often related to ingestion of contaminated food and water. With increasing evidence of antibiotic resistance in Shigella, researchers are trying to identify a potentially long term solution to prevent Shigellosis. Jun et al. (2016) assess the use of a bacteriophage as a way to control Shigella9. They use NGS to sequence the genome of Myoviridae phage, which showed efficient bacteriolytic activity against Shigella bacteria species. Bacteriophage can lyse a bacteria cell with high specificity without disrupting natural host microflora and phage therapy for humans has been of interest ever since the discovery of phages in 1915 and 1917. Jun et al. (2016) look into the possibility of using specific phages that can target and kill specific bacterial species and as a potential biocontrol agent and alternatives to antibiotics to control contaminated water and perhaps inhibit further antibiotic resistance9.
Shigella is a group of bacteria that cause Shigellosis, an infectious disease with a majority of Shigellosis cases occurring in developing countries. This bacterial pathogen is often related to ingestion of contaminated food and water. With increasing evidence of antibiotic resistance in Shigella, researchers are trying to identify a potentially long term solution to prevent Shigellosis. Jun et al. (2016) assess the use of a bacteriophage as a way to control Shigella9. They use NGS to sequence the genome of Myoviridae phage, which showed efficient bacteriolytic activity against Shigella bacteria species. Bacteriophage can lyse a bacteria cell with high specificity without disrupting natural host microflora and phage therapy for humans has been of interest ever since the discovery of phages in 1915 and 1917. Jun et al. (2016) look into the possibility of using specific phages that can target and kill specific bacterial species and as a potential biocontrol agent and alternatives to antibiotics to control contaminated water and perhaps inhibit further antibiotic resistance9.
Recently, several individuals in Wisconsin were infected and died due to bacterial contamination from a single source10. Although authorities have yet to identify the exact source of this outbreak, other, similar infections were narrowed down to contaminated tap water in a critical care unit10. The significance of this occurrence is the fact that these bacteria are resistant to antibiotics and disinfectants10. This is bad news, where we may potentially have a MDR bacterial outbreak. When it matters the most, it seems like we continue to utilize traditional quantitative techniques and fail to incorporate qualitative testing methods, which can be incorporated into current traditional testing practices. Technology such as PCR, qPCR, and next generation sequencing can potentially be used in addition to traditional testing methods to accurately identify the presence or absence of specific, even unculturable organisms.
W ater is a valuable resource and affects us all. Re-introducing antibiotics and antibiotic resistance genes into the environment may be tipping the scales for microbes to gain antibiotic resistance. Future research must focus on ways to improve wastewater treatment efficiency, to reduce the reintroduction of antibiotics, antibiotic resistance genes or MDR organisms back into the water supply or the environment. Further, closely testing and monitoring the water we drink not only for the number of microbes present, but to also understand the types of microbes present may help us control and prevent future outbreaks.
ater is a valuable resource and affects us all. Re-introducing antibiotics and antibiotic resistance genes into the environment may be tipping the scales for microbes to gain antibiotic resistance. Future research must focus on ways to improve wastewater treatment efficiency, to reduce the reintroduction of antibiotics, antibiotic resistance genes or MDR organisms back into the water supply or the environment. Further, closely testing and monitoring the water we drink not only for the number of microbes present, but to also understand the types of microbes present may help us control and prevent future outbreaks.
Let us know what you think in the comments section below about water testing in general, the use of available technology to improve the testing process, or how you feel about antibiotic resistance.
All products are for research use only. Not for use in diagnostic purposes
References:
1] Center for Disease Dynamics, Economics & Policy. 2015. State of the World’s Antibiotics, 2015. CDDEP: Washington, D.C.
2] Finkel SE, Kolter R: DNA as a nutrient: novel role for bacterial competence gene homologs. J Bacteriol 2001, 183(21):6288-6293.
3] Aydin S, Ince B, Ince O: Assessment of anaerobic bacterial diversity and its effects on anaerobic system stability and the occurrence of antibiotic resistance genes. Bioresour Technol 2016, 207:332-338.
4] Al-Jassim N, Ansari MI, Harb M, Hong PY: Removal of bacterial contaminants and antibiotic resistance genes by conventional wastewater treatment processes in Saudi Arabia: Is the treated wastewater safe to reuse for agricultural irrigation? Water Res 2015, 73:277-290.
5] Jutkina J, Rutgersson C, Flach CF, Joakim Larsson DG: An assay for determining minimal concentrations of antibiotics that drive horizontal transfer of resistance. Sci Total Environ 2016, 548-549:131-138.
6] Shaw JL, Monis P, Weyrich LS, Sawade E, Drikas M, Cooper AJ: Using Amplicon Sequencing To Characterize and Monitor Bacterial Diversity in Drinking Water Distribution Systems. Appl Environ Microbiol 2015, 81(18):6463-6473.
7] Graham RM, Doyle CJ, Jennison AV: Epidemiological typing of Neisseria gonorrhoeae and detection of markers associated with antimicrobial resistance directly from urine samples using next generation sequencing. Sex Transm Infect 2016.
8] Francy, D.S., Bushon, R.N., Brady, A.M.G., Bertke, E.E., Kephart, C.M., Likirdopulos, C.A., Mailot, B.E., Schaefer, F.W., III, and Lindquist, H.D.A., Performance of traditional and molecular methods for detecting biological agents in drinking water: U.S. Geological Survey Scientific Investigations Report 2009–5097, 17 p.
9] Jun JW, Giri SS, Kim HJ, Yun SK, Chi C, Chai JY, Lee BC, Park SC: Bacteriophage application to control the contaminated water with Shigella. Sci Rep 2016, 6:22636.
10] Tatera K, “People in Wisconsin Are Dying From a Mysterious Infection That Has Experts Stumped.” The Science Explorer. N.p., 22 Mar. 2016. Web. 29 Mar. 2016.
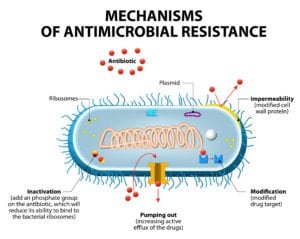
The Antibiotic resistance is a dirct result of overduse and abuse of the antibiotics in the food chain, including our water supplies. Water the most used and abused resource, with little care in management. The current water treatment technology is inefficianet at removing many pollutants such as oils, endocrine disrupters, pathogens and antibiotics.
Developping solutions to control or remove such pollutants is vital pr prtoect our water supplies, our environment and ultimately our health.
Yes, antibiotic resistance is an issue am very passionate about. We cede more ground to the MDR microorganisms everyday through: improper or excessive dosages: Use of antibiotics in agriculture and now it seems from waste water. Do you think there is a way we can research on how to destroy the genes in water treatment facilities before being released, as the traditional UV rays and other methods seem not to be working?
Hi Farah,
You are right, the global misuse and over prescription of antibiotics has played a major role in giving rise to MDROs. Other than ultrafiltration and osmosis, I am not aware of other techniques that can be applied on such a large scale in a cost-effective manner. More research is certainly required to help us remove these particles from our water supply.
One way to deal with the issue is to monitor the use and prescription of antibiotics. If we reduce the unnecessary use of heavy dose antibiotics, especially when not needed, it may help reduce the amount of antibiotics that make their way into the environment. Although this is a small step, it is better than not doing anything at all.
Technology today has certainly brought attention to something we weren’t really aware of: from evolution of MDROs, to understanding the effieciency of water treatment plants, to how (much) antibiotics in the water allow for transfer of genetic material.
With the use of bacteriophage to reduce infection are we not liberating more genetic material into the water. What about the use of ultra filtration to remove genetic particulate? I think that more work is needed to remove these antibiotics, pharmaceuticals, etc,. from the water our most precious resource.
Hi Heidi,
Yes you are right, we certainly may be introducing more genetic material into the water, but with fewer antibiotics in the environment, microbes may be less likely to gain resistance. Also bacteriophages allow us to target specific bacteria, whereas antibiotics tend to kill the good and the bad microbes in an ecosystem.
Ultra filtration and osmosis are certainly options, but I think the problem with that is the cost and energy it would require to remove all the nucleic acid particles and antibiotics, making water treatment a very expensive process. A process where more energy and resources are required to treat water, something like this cannot be done for large scale water treatment and still may not be 100% effective.
More research and novel ways to remove antibiotics and genetic material from the water are certainly required as the element of life becomes scarce.
Anyone think about HOW antibiotics are administered? Investigate that line of thought
Antibiotic resistance is a serious issue..and of course new technology has to be adminstered to treat water, but the challenge is being capable of using these methods in less developed countries that massively contribute to this high percent of contamination and death..
Very informative article you are sharing through your blog.
Follow Meddco Healthcare https://www.meddco.com for affordable healthcare packages available at your nearest hospital.
Thank You. Keep Sharing
If I can write like you, then I would be very happy, but where is my luck like this, really people like you are an example for the world. You have written this comment with great beauty, I am really glad I thank you from my heart.
Thank you very much for writing a great post. If there is anything in my life that I think is very important and I should understand it, I always try to understand it and I have seen your post. Saw this and liked this post a lot, so I am complimenting your post thoughtfully.
Thanks for giving us useful information about the Health, And do You know what the best 12 natural antibiotic?
This is the best and amazing website i love that
Sanganipackaging provide Packaging Material in Jaipur at best price. We provide best Packaging manufacturer, suppliers, wholesalers and dealers. We are best Packaging material manufacturer in Jaipur.
Ketovictoria is the world leader in Low Carb and Ketogenic Diet products. Execute a healthy Keto diet without changing your existing behaviour with our products. We provide many types Keto Products in India Your can visit our website and purchase online.
The fact of the matter is that Antibiotics are every where in today ground water. But another vital question is that “Is it only Antibiotics or lot of other things.
Factory chemicals. Sewerage, Medical Waste, industrial waste every thing is reaching our ocean. At the end it is damaging our bio diversity.
We stop industries then we loose consumerism and rich will become poor. So what is the solutions. Only one soluton is there. We will have to have local disposable unites in each pin codes.
The fact of the matter is that Antibiotics are every where in today ground water. But another vital question is that “Is it only Antibiotics or lot of other things.
Factory chemicals. Sewerage, Medical Waste, industrial waste every thing is reaching our ocean. At the end it is damaging our bio diversity.
We stop industries then we loose consumerism and rich will become poor. So what is the solutions. Only one soluton is there. We will have to have local disposable unites in each pin codes.
Hiiii
Hiii
Hiiii
Thanks for giving us useful information about the Health
Indocare Diagnostics & Pathology Lab is known for affordable health checkup. Full body checkup packages start from 599 INR. Full body checkup includes all essential tests. Assured top full body checkup at Book now & get best discount.Call us :7745887755
Very Nice post thank you for sharing us.