Genomic science has come a long way since the days of the Human Genome Project. The years have seen the development of newer, faster tools than the HGP could employ, and they now fill laboratories around the world, making genome sequencing an increasingly ordinary, even trivial task.
Detecting inherited chromosome anomalies more effectively
One of the most important steps in this advancement is the advent of microarrays, which enable the study of numerous gene targets and variants in a single assay. Microarrays make genomic analysis particularly simple compared to large-scale sequencing technologies, and array technology and methodology continues to improve both resolution and usability.
What is array competitive genomic hybridization?
One of the most common forms of microarray is array competitive genomic hybridization (aCGH). In an aCGH workflow, separate test and normal/control samples are each labeled with a different fluorescent dye. These samples are blended in a 1:1 mixture and then hybridized to the array. This allows the reader to compare the fluorescence of each dye, and the test-to-control fluorescence ratio reveals the differences between the test and control samples. CGH microarrays usually contain hundreds of thousands of probes, each aligning with a different locus on the target genome. This differs from single nucleotide polymorphism (SNP) arrays. In a SNP array, no control sample is necessary. The test DNA is labeled with a fluorescent dye and the absolute fluorescence bound to each probe on the microarray is measured, using a separate reference sample for standardization.
What is the difference between SNP arrays and array competitive genomic hybridization?
At the practical level, one of the biggest differences between these two methods is that SNP arrays are more effective at detecting a wide variety of structural abnormalities. SNP arrays typically contain both SNP and aCGH probes, combining the strengths of the two approaches to achieve greater results than aCGH can alone. For example, both of these methods can detect copy number variation (CNV), which is critical for studying many genetic conditions and abnormalities, but aCGH is not effective at detecting long continuous stretches of homozygosity (LCSH), reading this situation as a normal diploid. Long continuous stretches of homozygosity can arise in several ways, including parents who are related to one another, uniparental heterodisomy, uniparental isodisomy, and more, and all of these situations have distinctive SNP signatures but are difficult or impossible to detect via aCGH.
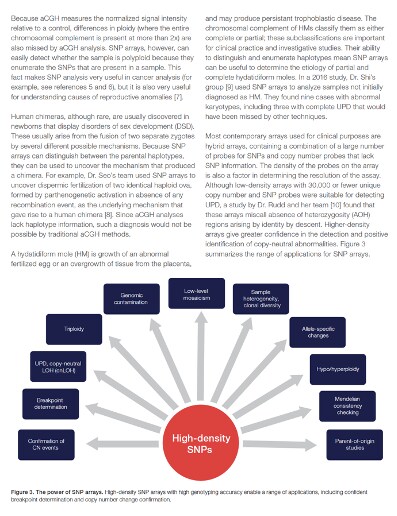
Similarly, SNP arrays are more effective than aCGH at detecting genomic contamination, sample heterogeneity, clonal diversity, allele-specific changes, and a wide variety of other small but important differences that aCGH cannot easily reveal. This makes SNP arrays particularly effective at detecting recessive conditions that run in families, imprinting disorders mediated by epigenetic differences, and neurodevelopmental disorders, all of which can be caused by the sorts of differences that SNP arrays are primed to identify.
SNP arrays are also capable of resolving unusual ploidy levels, in which one or more additional complete sets of chromosomes are present, and human chimeras, in which fusion of zygotes leads to different genotypes in different parts of a single person. Both of these situations can confound aCGH, due to the method’s use of normalized signal intensity relative to a control and inability to consistently distinguish between parental genotypes. Ploidy changes and chimerism are both important to understand: the former contributes to many cancers and the latter can complicate genotyping for the purposes of criminal investigation and transplant matching.
These differences in detection ability are not merely theoretical. SNP arrays’ ability to detect differences that aCGH cannot affects diagnostic outcomes. Shin et al. used SNP arrays to determine the mechanism underlying a human chimera—in this case, two identical ova fertilized by different sperm that then fused together.1 Similarly, De Noronha et al. were able to identify the underlying genetic causes of inherited deafness in a large Brazilian family, connecting it to large regions of homozygosity shared between various family members.2 A case of breathing difficulties and sinus invertus from a consanguineous family proved impossible to accurately diagnose until Edwards et al. used SNP arrays to identify long regions of homozygosity, leading to the recognition of an inherited mutation in LRRC6 and a diagnosis of primary ciliary dyskinesia (PCD), a serious lung condition.3 All of these conditions’ genetic signature were not detectable by aCGH and required SNP arrays to recognize and process, and they are far from the only such conditions.
As the costs of health care systems continue to be highly scrutinized, it is important to maximize the impact of every dollar spent. SNP arrays offer higher discovery yield—that is, more ability to detect abnormalities in each test—than other genotyping technologies, because they contain both CNV and SNP probes while being far less expensive and faster than whole-genome sequencing. SNP arrays are particularly ideal in situations in which cell culture has failed, which are below the resolution threshold for karyotyping, or in which maternal cell contamination complicates other methods.4 SNP arrays are also more able to extract usable information from degraded samples, such as formalin-fixed, paraffin-embedded (FFPE) samples used as references in cases of recurrent pregnancy loss.4 The greater diagnostic effectiveness of SNP arrays makes them ultimately more cost-effective than seemingly cheaper alternatives.5,6 For these reasons, SNP arrays often make an ideal first test that both yields extensive data and can suggest which other tests might most effectively yield more, and this approach is recommended by the American College of Medical Genetics and American College of Obstetricians and Gynecologists.
Clear diagnoses lead to fewer false starts in treatment, better health outcomes and lower healthcare costs. SNP arrays represent a sizable improvement over aCGH and other genomics technologies when it comes to drawing accurate conclusions from difficult genetic material, not least because they combine SNP and CNV probes into a single array. More widespread adoption of SNP arrays will illuminate many situations that currently remain mysterious and improve quality of life for large numbers of people.
For more information on prenatal and postnatal genetic testing, visit our educational page here
References
- Shin, S.Y., Yoo, H.-W., Lee, B.H., Kim, K.S., and Seo, E.-J. (2012) “Identification of the mechanism underlying a human chimera by SNP array analysis,” Am. J. Med. Genet. A. 158A, pp. 2119–2123.
- De Noronha, T.R., and De Lourdes Chauffaille, M. (2018) “Multiple long runs of homozygosity detected by SNP array: Offspring of consanguineous parents and his siblings,” Adv. Cytol. Pathol., Volume 3.
- Edwards, M., et al. (2016) “Syndrome diagnosis with single-nucleotide polymorphism (SNP) microarray,” J. Paediatr. Child Health, 52, pp. 85–89.
- Sahoo, T., et al. (2017) “Comprehensive genetic analysis of pregnancy loss by chromosomal microarrays: outcomes, benefits, and challenges.” Genet. Med. Off. J. Am. Coll. Med. Genet., 19, pp. 83–89.
- Sinkey, R.G., and Odibo, A.O. (2016) “Cost-effectiveness of old and new technologies for aneuploidy screening,” Clin. Lab. Med., 36, pp. 237–248.
- D’Amours, G., et al. (2014) “SNP arrays: Comparing diagnostic yields for four platforms in children with developmental delay,” BMC Med. Genomics, 7, p. 70.




Leave a Reply