Search
Microsatellite Marker Analysis

Microsatellite markers are co-dominant, polymorphic DNA loci containing repeated nucleotide sequences, typically with 2 to 10 nucleotides per repeated unit. The number of nucleotides in the repeated unit is the same for the majority of the repeats within an individual microsatellite locus, but the number of repeats for a specific locus may differ, resulting in alleles of varying length, which can be analyzed with fragment analysis by capillary electrophoresis. Because microsatellite markers are subject to Mendelian inheritance, analysis of length variation is a widely accepted tool for applications such as microsatellite instability (MSI) analysis.
Microsatellite instability (MSI) analysis
MSI is a form of genomic instability resulting in the accumulation of insertions or deletions (indels) in microsatellites during replication due to a dysfunctional mismatch repair (MMR) protein. MMR proteins are responsible for correcting errors made by DNA polymerase during replication. They do so by recognizing a temporary insertion-deletion loop that is created when DNA polymerase slips. Cells with a dysfunctional MMR protein accumulate errors when the loop resulting in frameshift mutations (indels), leading to the appearance of novel alleles at microsatellite loci, which can be easily identified via fragment analysis.
In addition to screening for cancers, MSI has recently been found to be predictive of response to immunotherapies. Tumors with defective MMR proteins often have somatic cells with mutations that produce novel proteins that can be immunogenic. Such tumors trigger an immune response and display susceptibility to immunotherapies.
MSI analysis involves comparing allelic profiles of specific microsatellite markers in normal and disease samples.

NEW TrueMark MSI Assay for microsatellite instability analysis
Experience more efficient and comprehensive MSI testing with automated calling, no normal controls required, and expanded 13 MSI marker panel.
Other applications
Chimerism is a term used to describe the occurrence of genetically distinct cell types in a single organism, which can result from transfusion or transplantation or can be inherited (e.g., in plants). In the case of a bone marrow transplant, the relative amounts of donor and recipient cells can be used to determine if engraftment has taken place.
Chimerism can be monitored via sensitive PCR-based methods and analysis of microsatellite markers. The sensitivity, improved data quality, and precision of our 3500 Series genetic analyzers, combined with our five-color chemistry and pre-optimized run conditions, create the ideal solution for chimerism analysis.
Short tandem repeats (STRs) are repeated DNA motifs consisting of 2 to 13 nucleotides. STR analysis measures the exact number of repeated units and compares specific loci on DNA from two or more samples. Uses for STR analysis include cell line authentication, human biobank sample matching, and ex vivo cell treatment tracking.
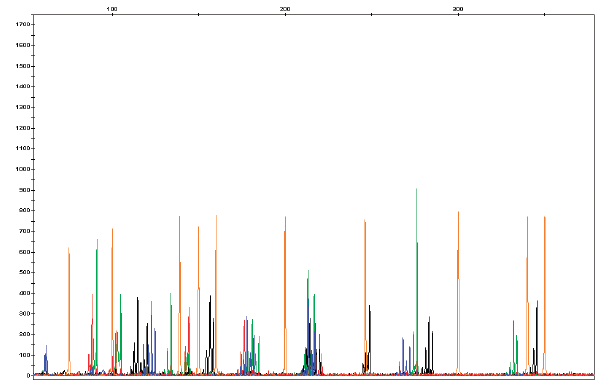
Inter-simple sequence repeat (ISSR) analysis is a technique commonly used for DNA fingerprinting, mostly in plant species, based on variation found in the regions between microsatellites. Since an ISSR may be a conserved or nonconserved region, this technique is not useful for distinguishing individuals, but has a wide range of research applications, including the characterization of genetic relatedness among populations, genetic fingerprinting, gene tagging, detection of clonal variation, cultivar identification, phylogenetic analysis, detection of genomic instability, and assessment of hybridization.
ISSR-PCR uses a single fluorescently labeled primer to target and amplify the region between identical microsatellites. The result is a mix of short, amplified DNA strands that vary widely in length.
Chimerism is a term used to describe the occurrence of genetically distinct cell types in a single organism, which can result from transfusion or transplantation or can be inherited (e.g., in plants). In the case of a bone marrow transplant, the relative amounts of donor and recipient cells can be used to determine if engraftment has taken place.
Chimerism can be monitored via sensitive PCR-based methods and analysis of microsatellite markers. The sensitivity, improved data quality, and precision of our 3500 Series genetic analyzers, combined with our five-color chemistry and pre-optimized run conditions, create the ideal solution for chimerism analysis.
Short tandem repeats (STRs) are repeated DNA motifs consisting of 2 to 13 nucleotides. STR analysis measures the exact number of repeated units and compares specific loci on DNA from two or more samples. Uses for STR analysis include cell line authentication, human biobank sample matching, and ex vivo cell treatment tracking.

Inter-simple sequence repeat (ISSR) analysis is a technique commonly used for DNA fingerprinting, mostly in plant species, based on variation found in the regions between microsatellites. Since an ISSR may be a conserved or nonconserved region, this technique is not useful for distinguishing individuals, but has a wide range of research applications, including the characterization of genetic relatedness among populations, genetic fingerprinting, gene tagging, detection of clonal variation, cultivar identification, phylogenetic analysis, detection of genomic instability, and assessment of hybridization.
ISSR-PCR uses a single fluorescently labeled primer to target and amplify the region between identical microsatellites. The result is a mix of short, amplified DNA strands that vary widely in length.
Microsatellite marker analysis workflow
Microsatellite marker analysis involves PCR amplification of the microsatellite loci using fluorescently labeled primers that flank the repeated sequence. The labeled PCR products are then analyzed by CE to separate the amplicons by size.


Build your knowledge
Get bite-sized answers to your everyday Sanger sequencing and fragment analysis questions
Learn about the history of sequencing and how to pick the right platform for your research needs


