Search
Axiom Microbiome Solution

The Axiom™ Microbiome Array enables the detection of all known microorganisms in a sample with species- and strain-level identification in a single, scalable assay. The proprietary photolithographic arrays help ensure fidelity and consistency across manufacturing batches, with designs available as long as required.
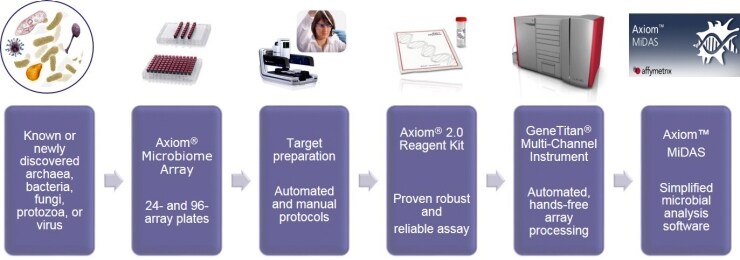
Utilizing the robust biochemistry of the Axiom™ 2.0 Assay coupled with manual and automated target preparation methods, this streamlined protocol enables consistent and high-quality results. A simplified, upstream reverse transcriptase reaction enables the detection of RNA virus genomes. Automated array processing is performed on GeneTitan™ Multi-Channel Instrument. The complete solution includes arrays, reagent kits, and free, easy-to-use microbiome analysis software.
Axiom™ Microbial Detection Analysis Software (MiDAS) provides a streamlined solution for the prediction of target identities in Axiom Microbiome Array data from an unknown sample. Features include complete analysis to strain-level, direct access to external NCBI databases, lists of microbial targets most likely present in each sample, and summaries of microbial content in both table and graphical formats to detect and profile the genetic composition of all microbes in a sample.
The Affymetrix Research Service Laboratory (ARSL) provides high-quality and economical processing of genomic DNA and reverse transcribed complimentary DNA from microbial samples. Complete analysis using Axiom MiDAS is included for detailed enumeration of microbes in each sample.
Using Axiom assay biochemistry, the Axiom Microbiome Solution interrogates non-polymorphic sequences in both family-conserved and target-specific regions from NCBI database sequences. The Axiom Microbiome Array detects over 12,000 species including archaea, bacteria, fungi, protozoa, and viruses. The array content is sample type agnostic, suitable for applications in nutrigenomics, agrigenomics, and animal research and modeling.
Highlights
- Expert design for profiling human and animal microbiomes with a single assay
- Detects greater diversity and higher resolution than 16s rRNA gene sequencing
- Comprehensive coverage of over 11,000 organisms and five microbial domains: archaea, bacteria, fungi, protozoa, and viruses
- Allows identification to the species, strain and sequence level; RNA virus detection using cDNA template
- Flexible 24- and 96-array plate formats
- Sample type agnostic: applications in nutrigenomics, agrigenomics, and animal research
- Simplified microbial detection analysis
Applications
- Assessment of animal gut microbial communities for feed optimization6
- Ascertainment of livestock animal health
- Microbial profiling for pathogen detection
- Disease prevention research
- Vivarium screening (monitoring for environmental and animal colony health)
- Evaluation of microbial status and response to treatment
- Evaluation of microbiota for soil productivity
- Linkage studies (impact of human gut microbiota on disease state)
- High-resolution profiling of probiotic and prebiotic mixtures
Resources
For Research Use Only. Not for use in diagnostic procedures.

