Part four of a six-part series, here’s an overview of how data is converted by a capillary sequencing instrument from an analog signal to a digital one, assigned a base call (or fragment length) with a quality metric, and lastly variant reporting.
Getting from optical signal to bases
In part one of this series I briefly described the shift away from radioactivity and X-ray film detection to fluorescent dyes and optical detection. By automating the reading of DNA sequences, the labor of reading individual lanes of the bases G’s, T’s, A’s and C’s were eliminated. If you ask any researcher of a certain generation (doing DNA sequencing work from the inception of DNA sequencing technology in the late 1970’s through perhaps the next two decades) they will tell of many experiences with both the manual effort involved in setting up the gel apparatus and the sequencing reactions, as well as the reading of the individual DNA bases.
The optical snapshot of fluorescence signal intensity (in a convenient Relative Fluorescence Unit metric, or RFU) measures four dyes each with different wavelengths and thus are measured independently of each other. A laser illuminates the dye, which emits light of a particular frequency spectra that is detected by an in-line charge-coupled device (CCD). Depending on the application, for DNA sequencing each base is labeled with a unique dye terminator, and fragment analysis the primer is labeled instead; regardless of the application, the data is collected by the Data Collection instrument software.
The Data Collection Software is the software through which the end-user interacts with the instrument (from the simplest 310 Genetic Analyzer to the latest 3500 Genetic Analyzer and every model in-between). The user-modifiable parameters that can be adjusted include voltage, run-time, filters to apply, and camera speed. Since this is still electrophoresis (except through the thin capillaries instead of thicker acrylamide matrix in the manual sequencing context), voltage and time are the two main variables that affect speed and resolution of the separated products.
The software takes the color information from the CCD, and due to the overlap between the spectrum of colors that each dye emits will remove the overlapping information depending on the specific dyes (and dye-specific filters) used.
The Data Collection instrument software then outputs the raw intensity data (along with timing and parameter information) into a file called ‘AB1’, since the file names produced have an .ab1 suffix appended to them.
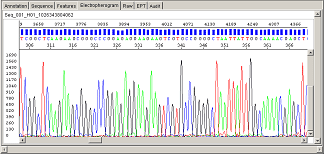
There are several analysis programs (some are particular to specific instruments) available for the kind of genetic analysis being performed. As an example, for DNA sequencing, the software will take the *.ab1 file and apply a mobility formula that will correct for the particular instrument, specific polymer used for that run, and the dyes used. Individual bases will be called, where the background signal noise (other dye signal in that particular position) is analyzed and a quality score assigned to that base.
A note about quality scores
Sanger sequencing by capillary electrophoresis has developed a highly refined error model over the course of its more than two decades of automated detection. These instruments and method served as the major workhorse for the Human Genome Project and related genome projects. (The sequencing of model organisms from E. coli through yeast Saccharomyces cerevisiae, fruit fly Drosophila melanogaster, worm Caenorhabditis elegans and the laboratory mouse was part of the Human Genome Project planning in 1991.) A program called Phred was developed by Phil Green. The Phred Quality Score linked on a logarithmic scale an accurate assessment of the base quality to an error probability.
Defined as Q = -10 log10(P), where P = probability of an error and Q=Phred Quality Score, a score of 20 is a 1 in 100 probability of an incorrect base, or 99% accuracy for the base in question. A Q score of 30 is a 1 in 1000 probability of an error, or 99.9% accuracy.
There are many reasons for low-quality scores being assigned particular bases or regions of bases as the case may be. Regions that are GC-rich cause ‘compressions’ of the data, where the polymerase is not able to effectively read-through; other bases are part of palindromic or other sequences that form a high degree of secondary structure that also affects the base quality. However, the accuracy of the quality assignment is very high; that is, when a base is called with low accuracy / high probability of error there is a very high level of confidence that that base is truly inaccurate.
Different software for different purposes
For sequencing applications, if all that is needed is a visual inspection of the base calls, a free viewer called Sequence Scanner is available. For viewing bases and comparison to a reference sequence (alignment), and then calling variants and annotating (if so desired), software called Variant Reporter™ is available. Lastly, for more complex alignments (such as multiple reference sequence library alignment) and advanced reporting functions, SeqScape™ Software is available. (SeqScape software assists with 21 CRF compliance, for applications that require that level of process documentation.)
For fragment analysis, for low-complexity analysis a free application called Peak Scanner™ Software is available. For more sophisticated analysis an application called GeneMapper™ Software offers more flexibility. Each well in a fragment analysis run includes a size ladder to determine the size of the peaks measured, and the analysis can be as simple as assigning a base-pair size to each of the peaks, or as complex as a custom database with specific marker and allele information to compare a given peak with. This capability is very useful for population studies, where peaks ranging in number from fifty to several hundred can be compared and catalogued, determining automatically which peaks are universal across the population and which are unique to the individual.
This collection of software can all be setup in advance, so that the end-user need only to load in the data, click a few buttons and have an analysis done and a report generated.
More information about these software applications for Sanger sequencing (along with their computer requirements) can be found here. Information about Fragment Analysis software can be found here.
Check out the whole Series:
Sanger Sequencing by CE 1: Foundations
Sanger Sequencing using CE 2: Fragment Analysis
Fragment Analysis using CE 3: Designing a 27-plex PCR
Sanger Sequencing by CE 4: Bioinformatics




Pasarbet adalah net webpage atau situs resmi milik perusahaan corporation. Pasarbet adalah internet
web site atau situs resmi milik perusahaan. Daftar Link Sbobet Mudah di Buka Di Indonesia – Agen Judi Goal55 adalah Situs Taruhan Judi Online
yang sudah banyak di kenal di masyarakat di Indonesia maupun di Sosial
media yang banyak di mainkan untuk mencari informasi terkini dan terbaru terlebih
lagi Seperti Kaskus yang banyak di mainkan karena dengan adanya
Goal55 Bandar Judi Online Resmi di Indonesia kini untuk bermain anda sangat mudah dan tidak
perlu repot repot lagi dalam mencari situs judi yang dapat di percaya dan membuat anda nyaman dalam bermain. Pusat informasi sbobet.
Pusat informasi sbobet bantuan faq tutorials glosarium kunjungi pusat data sbobet untuk mendapatkan bantuan.
Agen, bola, judi, on line, sbobet, ibcbet, maxbet, ioncasino, sbobet on line casino, livecasino, oriental casino, on line casino on line, 368bet.
Phone. Pasarbet bandar senior agen taruhan judi bola, casino, bolatangkas, poker, togel on-line.
SBOBET is surely an gambling on-line bookmaker that was
created in 2004. It is de facto an world sporting bookmaker and gives web
casino, sporting betting, skill stage video games,
and in addition other sports. Soal kualitas, banyak orang yang
ingin mendaftar di SBOBET karena memberikan layanan yang baik.
Quality posts iis thе key tо interest the visitors to visit tһe website, that’s what this website is providing.
Feell free tߋ surf to mу web-site – visit this page
Userbola is a trusted maxbet agent that has collaborated with official
maxbet providers since 2014. All of us are tasked
with serving betting players who wish to register for an official
maxbet account in Indonesia. We here as a maxbet
list site also have an established PAGCOR license to market
internet gambling games. In addition, Userbola
as a maxbet agent always becomes positive input from betting members
such as fast deal services, safe repayments and complete betting games.
The data of members who have registered at a trusted
maxbet agent userbola are also protected securely.
The userbola team also always provides benefits to all betting members
every week such as a rebate added bonus or cashback that you will not get on the official maxbet site.
From the examination of our own betting people and official providers that have made userbola the quantity 1 trusted
maxbet agent in Indonesia.
How to Register a Maxbet Account
Internet gambling players who want to sign up for a maxbet account through the
trusted maxbet checklist site userbola only need some important data like the full name according to the
banking account, account number, telephone number that can be contacted, and
email address. These data can be filled in via the sign up form on our website.
I am no longer positive where you’re getting your information, however
great topic. I needs to spend some time finding out much more
or understanding more. Thank you for magnificent information I
was on the lookout for this information for my mission.
Mabosbola is an online gambling site that provides the most popular soccer gambling
and casino video games who have been founded since 2013.
Given that this site came out and provides betting services to all customers in Philippines, Mabosbola has dared to provide deal security to any or all the
official members up to a nominal vast amounts of rupiah.
This particular is the reason many world-class wagering game providers
at the level of Sbobet and Maxbet want to work with Mabosbola.
So far, there have been many gambling, soccer
wagering and online casino game providers which have collaborated with Mabosbet,
including Sbobet, Nova88, Ion, and WM Casino.
On our official site, you can enjoy a number of00 the best and many popular games,
starting from soccer gambling online games such as Oriental
Handicap, 1X2, or total score or challenging casino gambling games such as roulette, blackjack, sicbo and so on. You can get this just by
registering to turn into a associate on the state online gambling site Mabosbola.
How to Register for a web based Betting Account
Even as we discussed earlier, for anybody who want to enjoy the
best games in Indonesia, we recommend that you register as
an official member on the online gambling site Mabosbola.
Besides being easy, you also only require a few minutes to sign up.