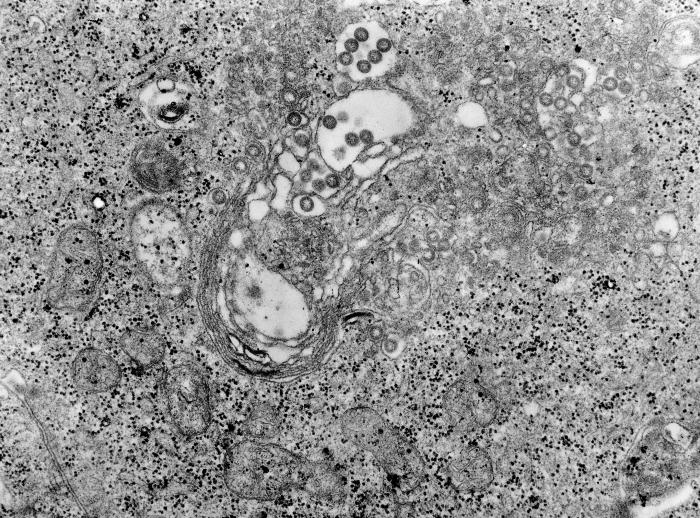
 Rift Valley fever is a potentially fatal disease affecting both humans and domestic animals. The disease is caused by Rift Valley fever virus (RVFV), of the family Bunyaviridae. Spreading via arthropods, symptoms in humans include fever, headache and liver abnormalities. In rare cases, humans may also encounter hemorrhagic fever syndrome, inflammation of the brain or even death. RVFV is especially fatal in livestock, with a nearly 100% abortion rate in pregnant animals and severe consequences in young livestock.
Rift Valley fever is a potentially fatal disease affecting both humans and domestic animals. The disease is caused by Rift Valley fever virus (RVFV), of the family Bunyaviridae. Spreading via arthropods, symptoms in humans include fever, headache and liver abnormalities. In rare cases, humans may also encounter hemorrhagic fever syndrome, inflammation of the brain or even death. RVFV is especially fatal in livestock, with a nearly 100% abortion rate in pregnant animals and severe consequences in young livestock.
RVFV’s potential use as a weapon of bioterrorism is great, as the virus is transmitted by mosquitos and can be genetically manipulated to become a more serious threat to humans.1 It is therefore important to identify proteins and pathways involved in the host response to RVFV, so that prevention and treatment options are available to the public. This summary describes recent findings identifying major host proteins involved in RVFV replication and transcription, along with therapeutics already in clinical trials.
To identify proteins essential for RVFV’s lifecycle, Nuss et al. (2014) observed virions purified from Vero cells, as compared to uninfected controls.2 By running virions and Vero cell controls via sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), the group isolated gel sections and performed liquid chromatography–tandem mass spectrometry (LC-MS/MS) using an LTQ Orbitrap XL hybrid ion trap-Orbitrap mass spectrometer (Thermo Scientific). From this experiment, the research team identified several host and endogenous RVFV proteins, including several glycoproteins and nucleocapsid proteins. Next, the team used an alternative approach to identify protein complexes involving both host and viral proteins. The approach included separating protein extracts using Blue Native PAGE, which uses non-denaturing conditions to separate protein complexes in their native states. The scientists then performed LC-MS/MS on the excised complexes. Because the cell line used in these experiments came from African green monkeys (Vero cells), which have a significant homology to the human proteome, Nuss et al. were able to utilize SEQUEST software (Thermo Scientific) and human protein databases to identify human host proteins interacting with RVFV.
After observing the complexes and individual proteins associated with RVFV infection in Vero cells, the group performed Western blots on RVFV virions to confirm the results of the LC-MS/MS experiments. The Western blots ascertained the association of seven host (Vero cell) proteins with RVFV virions. These host proteins included protein disulfide isomerase (PDI), heat shock proteins 70 and 90 (HSP70 and HSP90), glucose-regulated protein 78 (GRP-78), peroxiredoxin 6 (PDX 6), annexing A2, and annexing A4.
Bioinformatic studies identified roles that host proteins might play during RVFV infection—notably, host chaperones that had a major effect on the viral lifecycle. This finding was further supported by limiting protein expression using either siRNA gene silencing or chemical inhibitors to knock down host proteins and, in turn, functionally impact RVFV infection. Disrupting the expression of HSP70 and HSP90 (via siRNA or chemical inhibitors) especially demonstrated the importance of the proteins; infected cells survived the change but had major inhibition of RVFV. Interestingly, several of the chemical inhibitors for HSP90 are already in phase II clinical trials for cancer treatments. Finally, Nuss et al. presented evidence that these heat shock proteins play an important role during the post-entry replication/transcription phase of the RVFV lifecycle.
References
1. Flick, R., and Mandell, R.B. (2011) “Rift Valley fever virus: A real bioterror threat,” Journal of Bioterrorism & Biodefense, 2 (pp. 108–12).
2. Nuss, J., et al. (2014) “Multi-faceted proteomic characterization of host protein complement of Rift Valley fever virus virions and identification of specific heat shock proteins, including HSP90, as important viral host factors,” PLOS ONE, 9(5) (p. e93483), doi: 10.1371/journal.pone.0093483.
Post Author: Rebecca Easley.



Leave a Reply