Go Digital PCR: Pt 6 – Emerging Analyses is the final segment in our six-part series introducing you to digital PCR (dPCR) and now, how it aids absolute quantitation and rare sequence detection in lesser used applications.
dPCR helps optimize assay performance to enable additional sensitivity, precision, and absolute quantification using a wide range of widely used less widely used applications that include low level pathogen detection and quantification, to get an absolute pathogen count, and genetically modified organism (GMO) detection, with the sensitivity to find even a single errant GMO seed. dPCR is also quite useful in the absolute quantification of reference standards, when one doesn’t yet exist for your real-time PCR (qPCR) experiment.
The QuantStudio™ 3D Digital PCR System, with follow-up analyses using QuantStudio™ 3D AnalysisSuite™ Cloud Software, an easy-to-use data analysis tool with an intuitive user interface that allows multiple applications, including pathogen, viral load, and others.
The QuantStudio™ 3D Digital PCR System is capable of detecting both TaqMan® and SYBR™ chemistries. TaqMan® chemistry minimizes false positives in quantitative PCR experiments and eliminates the need for melt curve analysis. Over 8 million predesigned TaqMan® Assays are available to choose from when performing dPCR using the QuantStudio™ 3D Digital PCR System.
Absolute answers in just hours with QuantStudio™ 3D Digital PCR System
The QuantStudio™ 3D Digital PCR System is a simple and affordable dPCR platform for enabling nucleic acid detection and absolute quantification of rare alleles with the increased precision and sensitivity – a single molecule can be amplified a million-fold or more.
The QuantStudio™ 3D Digital PCR System leverages high-density nanofluidic chip technology to partition a sample into as many as 20,000 independent reaction wells. Comprised of consistently sized wells etched in a solid silicon substrate, the QuantStudio™ 3D Digital PCR Chip enables straightforward and robust sample partitioning that allows thousands of data points to be analyzed per run. The highly controlled sample loading and low reaction dropout combine to help ensure higher precision for absolute quantification in just a matter of hours.
The QuantStudio™ 3D Digital PCR System makes a data quality assessment for each calculation that determines absolute number copies/μL. Data considered marginal or failing are flagged for further review in secondary analysis using QuantStudio™ 3D AnalysisSuite™ Cloud Software.
Low-level pathogen detection and quantification
Detecting and quantifying low-level pathogens at the earliest possible opportunity is a key factor in reducing their impact in situations where bacterial or viral pathogens cause human illnesses, typically through contaminated food or water supplies.
While real-time qPCR can be a useful tool for pathogen detection, it remains sensitive to PCR inhibitors present in the crude samples used in food and water testing. Inhibitors do not easily affect dPCR because the inhibitors will be diluted in the partitioning, enabling the precise quantitation in situations where PCR efficiency is compromised. dPCR also enables an absolute quantification of pathogenic sequences, helping to eliminate the need for a reference material or a standard curve.
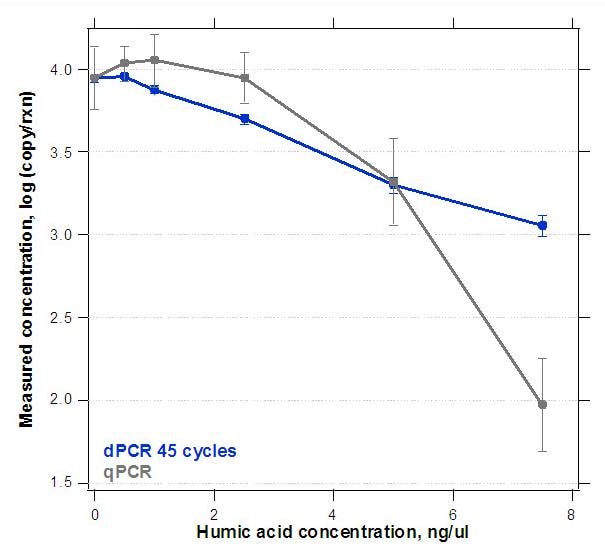
In this example of pathogen detection, a water sample was mock-contaminated with humic acid, a common environmental PCR inhibitor. The qPCR line (gray) shows drastic real-time PCR inhibition, as compared to the measurement of pathogen concentration by dPCR.
CAPTION: Deborah Grove, Director of Genomics Core Facility at Penn State Huck Institute of the Life Sciences, demonstrates key applications in low-level pathogen detection using the QuantStudio™ 3D Digital PCR System. The sensitive and absolute quantification capabilities of dPCR were used to detect a bacteria affecting Penn State campus elm trees as well as quantifying standards used in RNA virus real-time qPCR assays.
Sensitive GMO detection
Understanding the genetic complexity of plant phenotypes is no easy feat. The proliferation of GMO plants and calls for accurate food labeling requirements worldwide further compound the issue and increase the need for highly sensitive GMO detection.
Real-time qPCR has long been widely used for measuring plant gene expression. However as the concentration of target genes decrease amidst the presence of inhibitors, real-time qPCR quantification of plant mutations and detection of GMO has been challenging.
dPCR is ideal for GMO detection and measuring plant mutations because it allows you to decipher the expression of wild type versus mutant genes, helping to measure the absolute quantity of targets – specific alleles/SNPs – in a sample and detect new rare mutations. Again, dPCR requires no reference curve and also offers a high tolerance for many PCR inhibitors routinely encountered in plant samples.
Precision differential gene expression
Real-time qPCR is commonly used to detect differential gene expression but is generally limited to those that vary two-fold or more. The use of a reference or house keeping gene, e.g., actin, is typically required to express differential gene expression.
dPCR, with its ability to deliver highly precise measurements of ±10% or better, enable you to resolve changes of two-fold or less when performing differential gene expression. It can also eliminates the use of a reference gene to determine absolute quantification of a transcript.
Like real-time qPCR, dPCR requires the conversion of RNA to cDNA. Because conversion efficiency is important to experimental sensitivity, we offer a High-Capacity RNA-to-cDNA Kit, a streamlined reverse transcription kit that seamlessly integrates into your dPCR gene expression workflow.
Quantification precision comparison between dPCR (top) and real-time qPCR (bottom)

CAPTION: Sample 1 through 5 is a mixture of synthetic miRNA of hsa-miR-19b and hsa-miR-92 at different ratio: sample 1, 100%; sample 2, 95%; sample 3, 90%; sample 4, 75%; sample 5, 50%. After reverse transcription, cDNA was run with qPCR and dPCR with QuantStudio™ 3D Digital PCR System. Delta Ct of qPCR between hsa-miR-19b and hsa-miR-92 were reported for each sample. Relative quantitations for dPCR results were reported in percentile for each sample. dPCR with QuantStudio™ 3D Digital PCR System is able to discriminate a 5% difference between sample 1 and 2 (indicated by non-overlapping circles by Tukey-Kramer HSD test), while real-time qPCR was not able to discriminate even 10% difference between sample 1 and sample 3. Tukey-Kramer HSD test was done within JMP software with experiment replicates.
Absolute quantification of viral load
Determining the titer of a pathogenic virus relies on measuring the absolute quantification of molecules in a biological sample. But the determination of viral load poses unique challenges when using real-time qPCR, which is dependent on assay efficiency, instrument calibration metrics, and comparison to a reference sample of known concentration to convert Ct measurements for the unknown sample into an absolute concentration.
By contrast, dPCR is not dependent upon the amplification process cycle in which the reporter dye signal exceeds a Ct threshold. Instead, dPCR samples are partitioned into thousands of independent endpoint PCR reactions prior to amplification, and a reaction well is determined as either positive or negative for amplification of the viral sequence of interest. The negative wells are counted and converted to a concentration of target in the original sample.
The binary assignment of each reaction helps significantly minimizes the measurement’s dependency on assay efficiency and instrument calibration. This enables different laboratories to compare viral load measurement results in a standardized manner, without interference from external factors.
Absolute quantification of references and standards
Accurate genetic measurements often require comparison to reference samples and assay standards. For many organisms or applications, there’s often no suitable reference sample available. Generation of reference standards using conventional real-time qPCR requires you to consider how the reference sample will initially be calibrated, its long-term stability, and whether there is sufficient reference material for completion of all future studies. In addition, the lack of broadly adopted standards impacts comparison between laboratories.
Digital PCR precisely and accurately quantifies standards without the use of a standard curve
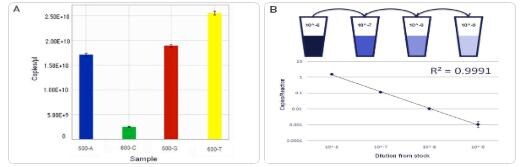
CAPTION: A: Four standards were measured in duplicate and results determined in absolute copies per microliter by dPCR. Tight error bars demonstrate high precision sample measurement. B: For sample 600-T, an additional 10-fold dilution series covering four logs of dilution was constructed. Copies per reaction for each dilution were calculated and demonstrate excellent correlation (0.9991), with extremely tight precision for each dilution.
dPCR does not require a reference sample or assay standard to enable absolute quantification which is determining the exact copy number of a nucleic acid target of interest. This is especially useful for calibrating reference samples and assay standards when none exist. Through direct copy number determination, digitally measured assay standards can enable you to compare results, with the assurance that your measurements are based on the same absolute baseline.
Conclusion
What is the key takeaway for this final segment of our six part series? dPCR helps optimize assay performance – to enable sensitivity, precision, and absolute quantification – for a wide range of non-mainstream applications, including low level pathogen detection and quantification, genetically modified organism (GMO) detection, and absolute quantification of reference standards,
when one doesn’t yet exist for your real-time PCR (qPCR) experiment.
Learn more about detection and quantification of emerging analyses
To learn more about how QuantStudio™ 3D Digital PCR System, and QuantStudio™ 3D AnalysisSuite™ Cloud Software enables absolute quantitation and rare sequence detection in less widely used applications.
Also available for your review is this recently published scientific study: S Patel, et al. Precise and Accurate Determination of MicroRNA Precursors by Digital PCR. J Biomol Tech. 2014 May; 25(Suppl): S27.
The study examines mounting evidence indicating that microRNAs are fundamental elements of almost every biological process. The researchers found that QuantStudio™ 3D dPCR technology offers significant improvement in precision and accuracy of measurement of a mir-9 primary precursor transcripts existing in a very diverse RNA world.
Read all of the Let’s Go Digital PCR Series:
Go Digital PCR: Pt 1 – Next-Gen Quantification
Go Digital PCR: Pt 2 – QuantStudio™ 3D
Go Digital PCR: Pt 3 – Copy Number Variation
Go Digital PCR: Pt 4 – NGS Libraries




Leave a Reply