Search
Recovering RNA From Small Samples RNAqueous™-Micro Kit
- Quantitative RNA recovery from 4 - 400,000 cells
- Generates concentrated RNA for immediate use in downstream applications
- Includes DNA-free™ technology to remove contaminating genomic DNA
Molecular analysis of gene expression from very small samples can be challenging. The RNA isolation method used for limited samples should provide for quantitative recovery of RNA that is intact, free from contaminants, and as highly concentrated as possible. Increasingly, expression analysis is being carried out on "micro-sized" samples, including samples obtained by Laser Capture Microdissection (LCM) and related microdissection techniques; needle biopsies; fine dissection; and samples comprised of low numbers of cultured cells. In these cases, it is desirable to recover the total RNA in a small volume so that the entire sample can be used in downstream applications, such as reverse transcription. Also, keeping the sample as concentrated as possible facilitates analysis and quantitation of the total RNA using sensitive microfluidic assays (e.g. Agilent 2100 bioanalyzer). Ambion's RNAqueous™-Micro Kit was developed to optimize recovery of highly concentrated total RNA from micro-sized samples.
Isolate Exceedingly Pure Total RNA in a Small Volume
The RNAqueous-Micro Kit begins with sample disruption in a chaotropic solution to lyse the cells and inactivate cellular nucleases. The cell lysate is loaded onto a glass fiber filter, washed and eluted. The filter cartridge provided in the RNAqueous-Micro Kit is only a few millimeters in diameter so that it can be saturated with a small volume of fluid. This allows the RNA to be eluted in a volume of only 20 µl. After elution, the RNA is treated with DNase I to eliminate genomic DNA contamination that can interfere with RT-PCR assays. The DNase is then removed using the
DNA-free™ Removal Reagent which removes the DNase I and buffer ions in a simple, quick step without organic extraction or heat inactivation. (Heat inactivation of DNase frequently causes divalent cation-mediated degradation of the RNA.) Using the RNAqueous-Micro Kit, the entire RNA isolation procedure, including DNase treatment, takes about 30 minutes.
High Quality RNA for qRT-PCR
To demonstrate the quality of RNA isolated by the RNAqueous-Micro Kit, RNA from 4 - 400,000 cells was recovered using the procedure and analyzed for integrity and concentration. Figure 1B shows an electropherogram of some of this total RNA. The RNA was intact. The RNA was then used in a one step qRT-PCR assay, and linear recovery of mRNA was shown over a range of 1 100,000 cell-equivalents (diluted from the 4 - 400,000 cells; Figure 1A). Minus RT reactions showed no amplification (data not shown).
Figure 2 shows an electropherogram and real-time PCR data generated from cells isolated by LCM and RNA purified using the RNAqueous-Micro Kit. The RNAqueous-Micro Kit contains sufficient reagents and filter cartridges to isolate total RNA free of DNA from 50 samples of 1 - 1 x 105 cells or up to 5 mg of tissue.
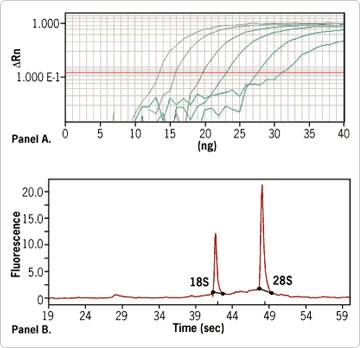
Figure 1. (A) Linear Recovery of RNA from Cells Over a 5 Log Range. RNAqueous™-Micro Kit was used to isolate total RNA from K562 cells diluted from 100,000 cell equivalents down to 1 cell equivalent. The RNA samples were then used for amplification of GAPDH by real time RT-PCR. (B)Electropherogram of RNA isolated from 10,000 K562 cells. RNA was run on an Agilent 2100 bioanalyzer. Total yield after DNase treatment and clean up was 360 ng with an 28S/18S rRNA ratio of 1.87.

Figure 2. (A) Use of LCM Samples for One Step RT-PCR of Neurogranin. RNA was isolated from nine mouse brain Laser Capture Microdissection samples using the RNAqueous™-Micro Kit. The 20 µl post isolation eluates were DNase treated (final volume 25 µl). A fifth of this treated sample was subjected to one-step RT-PCR for analysis of neurogranin (Ct of 18.44).
(B) Electropherogram of RNA Isolated from the Granule Cell Layer of Hippocampus (Mouse Brain). Cells were obtained through LCM and RNA was isolated with the RNAqueous™-Micro Kit. 5 µl was run and analyzed on an Agilent 2100 bioanalyzer. Total yield after DNase treatment and clean up was 120 ng with an 28S/18S rRNA ratio of 0.72.
Figure 2 shows an electropherogram and real-time PCR data generated from cells isolated by LCM and RNA purified using the RNAqueous-Micro Kit. The RNAqueous-Micro Kit contains sufficient reagents and filter cartridges to isolate total RNA free of DNA from 50 samples of 1 - 1 x 105 cells or up to 5 mg of tissue.

Figure 1. (A) Linear Recovery of RNA from Cells Over a 5 Log Range. RNAqueous™-Micro Kit was used to isolate total RNA from K562 cells diluted from 100,000 cell equivalents down to 1 cell equivalent. The RNA samples were then used for amplification of GAPDH by real time RT-PCR. (B)Electropherogram of RNA isolated from 10,000 K562 cells. RNA was run on an Agilent 2100 bioanalyzer. Total yield after DNase treatment and clean up was 360 ng with an 28S/18S rRNA ratio of 1.87.

Figure 2. (A) Use of LCM Samples for One Step RT-PCR of Neurogranin. RNA was isolated from nine mouse brain Laser Capture Microdissection samples using the RNAqueous™-Micro Kit. The 20 µl post isolation eluates were DNase treated (final volume 25 µl). A fifth of this treated sample was subjected to one-step RT-PCR for analysis of neurogranin (Ct of 18.44).
(B) Electropherogram of RNA Isolated from the Granule Cell Layer of Hippocampus (Mouse Brain). Cells were obtained through LCM and RNA was isolated with the RNAqueous™-Micro Kit. 5 µl was run and analyzed on an Agilent 2100 bioanalyzer. Total yield after DNase treatment and clean up was 120 ng with an 28S/18S rRNA ratio of 0.72.
Working With Laser Capture Microdissection Samples Laser Capture Microdissection (LCM) is a method for obtaining pure populations of cells from heterogeneous samples. LCM permits the selection and capture of cells, cell aggregates, and discrete morphological structures from thin tissue sections. For LCM, frozen and OCT embedded, or formalin-fixed and paraffin embedded tissue sections are processed, stained, and dehydrated, then scanned under an inverted microscope. Cells are visualized through a thermoplastic film which is attached to the bottom of an optically clear microfuge tube cap. A laser pulse, directed onto the target cells through the film, melts the film and allows it to flow onto the targeted area where it cools and bonds with the underlying cell(s). The film including the adhered cells or clusters is then lifted. Captured cells can then be used for nucleic acid studies, including SNP analysis, endpoint and real-time RT-PCR, and mRNA expression profiling.