Search
Assessing Gene Function with siRNA Libraries
Sets of siRNAs focused on a specific gene class, also called siRNA libraries, have the capacity to greatly increase the pace of pathway analysis and functional genomics. Here is an example of the power of Ambion's Silencer™ Kinase siRNA Library, as demonstrated through preliminary screening experiments carried out by Cenix BioScience GmbH. This highly validated library includes nearly 1800 individual siRNAs targeting 597 human kinases. In this experiment, 178 siRNAs from the library, targeting 178 different kinases expressed in HeLa cells, were each tested to determine their effects on cellular proliferation and mitotic index. This screen identified several kinase genes that are involved in the cell cycle, confirmed the identify of some kinases known to be involved, and identified a few kinases that had not previously been reported to be involved.
Experimental Design
The Silencer Kinase siRNA Library includes more than 600 validated siRNAs, which have been shown to reduce target mRNA levels by >70% when transfected into HeLa cells at a concentration of 100 nM. Cenix BioScience chose 178 of these validated siRNAs, targeting 178 different kinases, for their experiment. These siRNAs were individually transfected in triplicate at a final concentration of 100 nM into HeLa cells, plated at approximately 8000 cells per well in 96 well plates 24 hours prior to transfection. A validated siRNA targeting cyclin B1 was used as a positive control. Cyclin B1 is a well-studied cell cycle regulator known to play a key role in initiating mitosis, and therefore was expected to yield a clear decrease in mitotic index in this study. The Silencer Negative Control #1 siRNA (Ambion) was used as a "scrambled" sequence negative control.
Forty eight hours after transfection, cells were fixed and stained with DAPI to reveal chromatin, with antitubulin for microtubule distribution analysis, and with an antiphosphohistone H3 antibody to identify mitotic cells. The extent of cell proliferation was monitored by counting the number of cells in each well. The percentage of cells undergoing mitosis, or 'the mitotic index', was evaluated by fluorescence microscopy. Additionally, RNA was isolated from parallel samples and reverse transcribed to produce cDNA, and then target levels were measured by real-time PCR.
Verifying siRNA Efficacy
An important component of data analysis in any siRNA experiment is to monitor the extent of mRNA degradation, or 'knockdown', elicited by a particular siRNA. Real-time PCR is a rapid and sensitive technique that is ideally suited for this purpose. Figure 1 shows the target mRNA remaining 48 hours after siRNA transfection as monitored by real-time PCR. For each kinase siRNA, target mRNA levels were reduced by 70% or more as compared to the levels of mRNA obtained after transfection with the negative control siRNA.
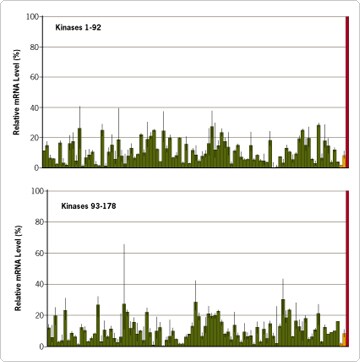
Figure 1. mRNA Silencing by 178 Kinase siRNAs from the Silencer™ Kinase siRNA Library. HeLa cells were plated at approximately 8000 cells per well in 96 well plates. Twenty-four hours later the cells were transfected in triplicate with each siRNA (100 nM). Forty-eight hours post-transfection, RNA was isolated, converted to cDNA, and analyzed by real-time PCR. Shown are the relative mRNA levels compared to cells transfected with a control scrambled siRNA (red bar, Silencer Negative Control #1 siRNA).
Effects on Cell Proliferation
Figure 2 demonstrates the effect of each siRNA on cell proliferation. In wells in which the negative control siRNA was transfected, there were approximately 1,250 cells/microscopic image after 48 hours (Figure 2). Inhibition of cyclin B1, which is known to be critical for initiation of mitosis, results in growth arrest. As expected, cell proliferation was dramatically inhibited by the siRNA targeting cyclin B1; there were <700 cells/microscopic image in wells transfected with this siRNA. Figure 2 also shows several kinase siRNAs that inhibit cell proliferation. This inhibition can result from a number of underlying causes that fall into three broad categories: effects causing cell necrosis, effects causing apoptosis, and effects causing cell cycle deregulation. Interestingly, a few kinase siRNAs appear to have a stimulatory effect on cell proliferation.
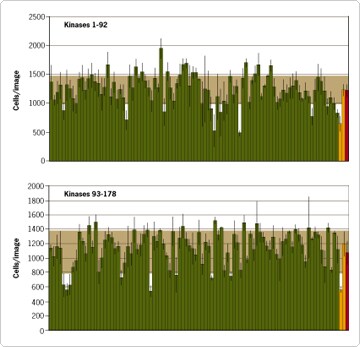
Figure 2. Changes in Cell Proliferation Induced by 178 Kinase siRNAs from the Silencer™ Kinase siRNA Library. HeLa cells were transfected with individual siRNAs targeting 178 different kinases as described in Figure 1. Forty-eight hours post-transfection cells were counted. Cell numbers per microscopic image are given.
Changes in Mitotic Index
One way to better understand a gene's role in cell cycle progression is to monitor the percent of cells undergoing mitosis at any given point in time, with and without siRNA treatment. For asynchronously growing cultures such as those used in these experiments, the mitotic index reflects the fraction of time that cells spend in mitosis versus the rest of the cell cycle. Thus, as shown in Figure 3, the ~2.5% mitotic index value observed for control cells transfected with a scrambled siRNA indicates that cells spent 2.5% of the cell cycle in mitosis.
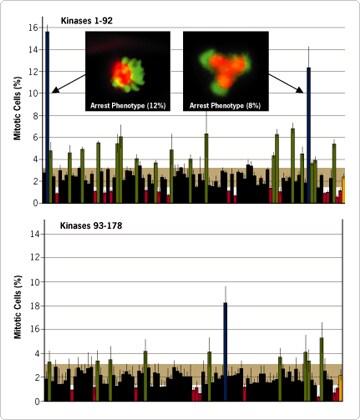
Figure 3. Mitotic Index of Cells Transfected with 178 siRNAs from the Silencer™ Kinase siRNA Library. HeLa cells were transfected with individual siRNAs targeting 178 different kinases as described in Figure 1. Forty-eight hours post-transfection mitotic index was measured. The orange bar shows the percent of negative control cells, which were transfected with a control scrambled siRNA, undergoing mitosis (mitotic index range of 2.1 to 2.3). The semitransparent brown rectangles highlight the 95th percentile range of normal mitotic index. The inset shows two interesting mitotic arrest phenotypes induced by siRNAs targeting two different kinases (green = chromatin; orange = tubulin).
Several siRNAs induced dramatic changes in the mitotic index when monitored 48 hours after siRNA transfection. Decreases in mitotic index reflect either a shortening of mitosis or a lengthening -- or 'arrest' -- of interphase. Conversely, increased mitotic index results from a lengthening of mitosis (usually an arrest) or a shortening of interphase. Both of these results were seen among the siRNAs tested. Two of the siRNAs increased the mitotic index more than five fold.
The mitotic arrest phenotypes were then examined by microscopic readout to determine their root causes. This was done by immunofluorescence staining of multiple markers, including tubulin to reveal microtubule distribution. This approach revealed one kinase siRNA that triggered a prometaphase-like arrest state (nearly 16% mitotic index, or ~7-fold higher than normal) wherein spindles were clearly bipolar and chromosomes were well condensed, but with no evidence of successful chromosome alignment (Top Panel, left inset). This likely corresponds to activation of the well-studied spindle assembly checkpoint, which monitors the correct, functional interactions between spindle microtubules and kinetochores in preparation for metaphase and subsequent anaphase onset. A second kinase siRNA that also generated cells in mitotic arrest induced the formation of aberrant spindles displaying either too many or too few apparent poles (e.g. Top Panel, right inset).
Putting It All Together
This screening experiment illustrates the power of higher content, multiparameter assays combined with libraries of effective siRNAs. This particular data set allows the researcher to go beyond a primary readout (affects on cell proliferation in this case) to efficiently and directly address underlying causes such as cell cycle deregulation at several levels with a single screening experiment. By comparing the cell proliferation and mitotic index data, one can relate the antiproliferative effects observed for several kinase-targeting siRNAs back to underlying effects on cell cycle regulation.
Silencing kinases necessary for passage of the cell through G1, S, and G2 phases of the cell cycle may be expected to lengthen interphase and therefore decrease the percentage of cells in mitosis. It will also decrease the overall cell proliferation rate. Several of the siRNAs investigated here induced this phenotype (Figure 4). Similarly, inhibition of kinases required for passage through mitosis would be expected to cause an increase in mitotic index and a decrease in cell proliferation. Several siRNAs also appeared to induce this phenotype.
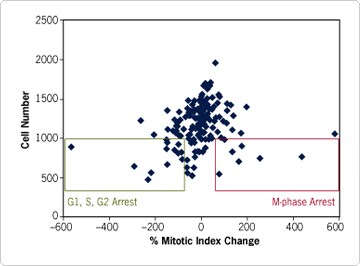
Cases where changes in cell proliferation do not correlate with aberrant mitotic index warrant follow-up analysis of necrosis and apoptosis as likely causes. With this in mind, Cenix has recently developed a multiplex assay monitoring cell proliferation, mitotic index, necrosis, and apoptosis all within one screening experiment. Using this and other basic and/or disease-focused RNAi assays, Cenix is further investigating the roles of kinases and other genes covered by Ambion's Silencer siRNA libraries.