Search
MELT™ Total Nucleic Acid Isolation System
As with many cancers, the molecular underpinnings of prostate cancer (PrCa) are poorly defined. Animal models, such as xenograft mice, offer an invaluable, in vivo context for studying both the etiology and progression of disease. Such models provide a path to cancer experimentation that can lend insights into the underlying biology and test the efficacy of new treatments. Recently, Ambion scientists began a collaboration with Drs. Jin-Rong Zhou and Sandra Gaston (Beth Israel Deaconess Medical Center, Harvard Medical School) and scientists at Illumina, Inc. to assess the global gene expression profiles in a mouse xenograft model for PrCa. An overview of their results is presented here.
Prostate Cancer
Prostate cancer (PrCa) is one of the most frequently diagnosed cancer, striking one out of every six men during his lifetime. Molecular analysis of disease progression in PrCa is critical for developing effective treatments for different patients (see sidebar, Prostate Cancer Cells and Hormone Responsiveness). Dr. Zhou, Dr. Gaston, and scientists from Ambion, Asuragen, and Illumina have designed experiments to examine temporal changes in global gene expression profiles and identify genes that are important during different stages of prostate cancer.
Xenograft Mouse Model
Approximately 2 million human prostate cancer LNCaP cells were injected into the ventral lobe of the prostate of several immuno-compromised (SCID) mice to seed the growth of a human tumor. Following expansion of the tumor, a subset of animals was castrated to eliminate the production of androgen hormones and induce the emergence of androgen-independent PrCa. Prostate tumors were harvested at intervals of 1-4 weeks. The collection of samples over this time frame provided a “snapshot” of the gene expression profile as the PrCa transitioned from an androgen-sensitive to an androgen-independent phenotype.
Preparing RNA from Tumors
Total RNA was extracted from 12 biological replicate samples at each time point using the MELT™ Total Nucleic Acid Isolation System (see the sidebar,
MELT™ Total Nucleic Acid Isolation System, for more information). RNA quality was assessed using the RNA integrity number (RIN), which is based on the entire electrophoretic trace of the RNA sample (Figure 1). Duplicate RNA samples (250 ng/ sample) from each prep were then amplified and labeled using Ambion’s
Illumina® TotalPrep™ RNA Amplification Kit following the recommended protocol. Yields of aRNA were typically 25 µg, with average lengths of ~1500 nt, as measured by the Agilent® 2100 bioanalyzer. Labeled RNA (850 ng) from a total of 12 biological replicates was hybridized to an Illumina Sentrix® HumanRef-8 BeadChip, containing 24,385 probes targeting human genes.
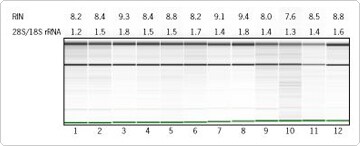
Figure 1. Integrity of RNA Isolated by MELT™ Total Nucleic Acid Isolation System. Total RNA (80–220 ng), isolated using the MELT Total Nucleic Acid Isolation System was run on an Agilent® 2100 bioanalyzer and is displayed here as a gel electropherogram. Samples 1 through 12 are the 12 biological replicates used in this research. RIN=RNA Integrity Number (10=high integrity, 1=low integrity)

Figure 1. Integrity of RNA Isolated by MELT™ Total Nucleic Acid Isolation System. Total RNA (80–220 ng), isolated using the MELT Total Nucleic Acid Isolation System was run on an Agilent® 2100 bioanalyzer and is displayed here as a gel electropherogram. Samples 1 through 12 are the 12 biological replicates used in this research. RIN=RNA Integrity Number (10=high integrity, 1=low integrity)
Microarray Analysis
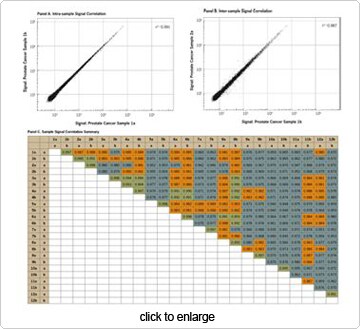
As shown in Figure 2, correlation of technical replicates (samples “a” and “b” for each number designate) was very high (mean R
2=0.996 ± 0.003). Correlations between biological replicates were generally ~0.98. However, this concordance decreased for samples representing more advanced tumor growth. This is not an unexpected result, since the accumulation of biological changes that accompany cancer progression can create additional variability in the gene expression profiles observed in more advanced prostate cancer.
Figure 2. Sample Signal Correlation by Illumina® Sentrix® Gene Expression Microarray. (A) Excellent signal correlation was observed between technical replicates. (B) Excellent signal correlation was generally observed between biological replicates, but decreased with advanced cancer progression. (C) This study contained 12 biological replicates, with technical replicates designated by an “a” or “b.” Green shading indicates inter-sample signal correlations above 0.990, orange shading indicates inter-sample signal correlations that are >0.980 and <0.990, and gray shading indicates inter-sample signal correlations that are <0.980.M
The array data were evaluated to identify genes with expression levels that varied at least 2-fold in androgen-independent vs. androgen-sensitive tumor tissue; nearly 50 genes met this criteria. The combination of the MELT Total Nucleic Acid Isolation System, Illumina TotalPrep RNA Amplification Kit , and Illumina Sentrix Array provided a powerful solution from sample-to-data to identify candidate RNA biomarkers of PrCa. A manuscript describing these results in full detail is in preparation.
Figure 2. Sample Signal Correlation by Illumina® Sentrix® Gene Expression Microarray. (A) Excellent signal correlation was observed between technical replicates. (B) Excellent signal correlation was generally observed between biological replicates, but decreased with advanced cancer progression. (C) This study contained 12 biological replicates, with technical replicates designated by an “a” or “b.” Green shading indicates inter-sample signal correlations above 0.990, orange shading indicates inter-sample signal correlations that are >0.980 and <0.990, and gray shading indicates inter-sample signal correlations that are <0.980.M
The array data were evaluated to identify genes with expression levels that varied at least 2-fold in androgen-independent vs. androgen-sensitive tumor tissue; nearly 50 genes met this criteria. The combination of the MELT Total Nucleic Acid Isolation System, Illumina TotalPrep RNA Amplification Kit , and Illumina Sentrix Array provided a powerful solution from sample-to-data to identify candidate RNA biomarkers of PrCa. A manuscript describing these results in full detail is in preparation.
Prostate Cancer Cells and Hormone Responsiveness
Initially, most prostate cancers express androgen receptor (AR) and are androgen sensitive, meaning that the growth and survival of these cells depends upon stimulation by testosterone or dihydrotestosterone. Anti-androgen therapies can be used to check the growth of hormone dependent prostate cancers, with cell death causing shrinkage of the tumor mass. However, over time the prostate cancer cells that are able to survive anti-androgen therapy will resume growth as an androgen-independent tumor. The emergence of androgen-independent growth is a sign of an aggressive prostate cancer with a poor prognosis.
MELT™ Total Nucleic Acid Isolation System
The MELT System (patent pending) uses a unique, hands-free, closed-tube enzymatic digestion to liquefy tissues and irreversibly destroy RNases. The RNA isolation component of this system uses MagMAX™ magnetic bead-based technology that is amenable to both manual and automated processing. The MELT System is designed to purify total RNA from ≤10 mg samples of fresh or frozen tissue and is recommended for use with most animal tissues that do not contain comparatively high levels of RNases and that are not extremely hard or fibrous. It is not compatible with tissues that have been stored in Ambion’s RNAlater® or RNAlater-ICE.
Because of the relatively high RNase activity in the tumor implant samples in this study, samples were processed in small batches (six at a time), and the MELT digestion time was rigorously controlled (10 min) to minimize variability and maximize RNA yield. All lysates were stored overnight at –20°C, and RNA was recovered the following day in batches of 12–16 samples.
Because of the relatively high RNase activity in the tumor implant samples in this study, samples were processed in small batches (six at a time), and the MELT digestion time was rigorously controlled (10 min) to minimize variability and maximize RNA yield. All lysates were stored overnight at –20°C, and RNA was recovered the following day in batches of 12–16 samples.