Search Thermo Fisher Scientific
Product Details
TA805328
Species Reactivity
Host/Isotype
Class
Type
Clone
Immunogen
Conjugate
Form
Concentration
Purification
Storage buffer
Contains
Storage conditions
Shipping conditions
Target Information
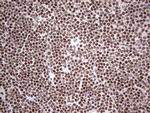
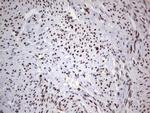
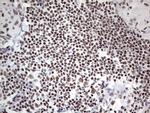
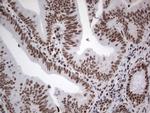
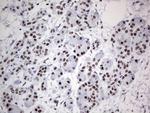
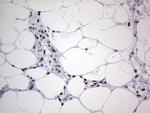
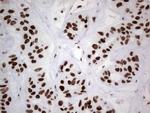
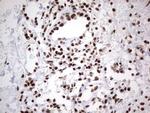
In the intact cell, DNA closely associates with histones and other nuclear proteins to form chromatin. The remodeling of chromatin is a critical component of transcriptional regulation and the acetylation of nucleosomal histones is a major source of this remodeling. Acetylation of lysine residues in the amino terminal tail domain of histone results in an allosteric change in the nucleosomal conformation and an increased accessibility to transcription factors by DNA. Several mammalian proteins function as nuclear histone acetylases, including GCN5, PCAF (p300/CBP-associated factor), p300/CBP, HAT1 and the TFIID subunit TAF II p250. Conversely, the deacetylation of histones is associated with transcriptional silencing. The histone deacetylases (HDAC) include HDAC1-9. HDAC9 and HDAC9a are two alternatively spliced isoforms of HDAC9. HDAC9a is 132 amino acids shorter than HDAC9, but both isoforms contain the HDAC catalytic domain, remain capable of deacetylase activity and repress myoctye enhancer-binding factor 2-mediated transcription. HDAC9 and HDAC9a are expressed in brain, skeletal muscle, kidney, placenta and pancreas.
For Research Use Only. Not for use in diagnostic procedures. Not for resale without express authorization.
References (0)
Bioinformatics
Protein Aliases: DKFZp779K1053; HD7; HD9; hdac-9; histone deacetylase 4/5-related protein; Histone deacetylase 7B; Histone deacetylase 9; Histone deacetylase-related protein; MEF-2 interacting transcription repressor (MITR) protein; MEF2-interacting transcription repressor MITR
Gene Aliases: HD7; HD7b; HD9; HDAC; HDAC7; HDAC7B; HDAC9; HDAC9B; HDAC9FL; HDRP; KIAA0744; MITR
UniProt ID: (Human) Q9UKV0
Entrez Gene ID: (Human) 9734
Molecular Function:
![]() histone modifying enzyme
histone modifying enzyme

Performance Guarantee
If an Invitrogen™ antibody doesn't perform as described on our website or datasheet,we'll replace the product at no cost to you, or provide you with a credit for a future purchase.*
Learn more
We're here to help
Get expert recommendations for common problems or connect directly with an on staff expert for technical assistance related to applications, equipment and general product use.
Contact tech support