Search
Mutation Detection Support—Troubleshooting
Having difficulties with your experiment?
We are dedicated to your success. Get back on track. View our expert recommendations for commonly encountered problem scenarios.
View the relevant questions below:
Beginning your experiment?
Visit our
Some mutant allele assays, designed to difficult target sequences, are expected to give low-level nonspecific amplification of wild type samples. Refer to the TaqMan® Mutation Detection Assay Index File, which provides the off-target amplification Ct values for each mutant allele assay, for more details. Another possibility is that the negative control wells have been contaminated. Check all reagents as needed.
Some mutant allele assays could potentially exhibit cross-reactivity with nontarget mutations that are found at the same or nearby genomic nucleotide positions as the target mutation. For example, an assay that detects a specific EGFR exon 19 mutation might also show some amplification signal with samples containing other, nontarget exon 19 mutations due to mismatch base pairing by the allele-specific primer. If a sample containing a nontarget mutation gives a positive result, it is likely to be caused by cross-reactivity. For the validated TaqMan® Mutation Detection Assay set, experiments were conducted to investigate and establish cross-reactivity patterns for mutant allele assays run with synthetic DNA templates containing mutations. No significant cross-reactivity (≥0.1%) was detected for the validated assay set.
Cross-reactivity studies have not been run for other mutant allele assays, but judging from the studies done with the validated assay set, mutant allele assays for point mutations in codons are expected to have no significant cross-reactivity with other point mutations in the same or nearby codons. Mutant allele assays designed to detect deletion mutations may have a low level of cross-reactivity with deletion mutations that are similar in sequence. As well, assays that detect single point mutations may detect related multiple-nucleotide mutations if the single point mutation is positioned at the first or last base of a multiple-nucleotide mutation. In general, if the on- and off-target Ct values differ enough (for example, ∆Ct of 6.7 would be 1% cross-reactivity), the activity is not a concern since you would still be able to differentiate between true hits.
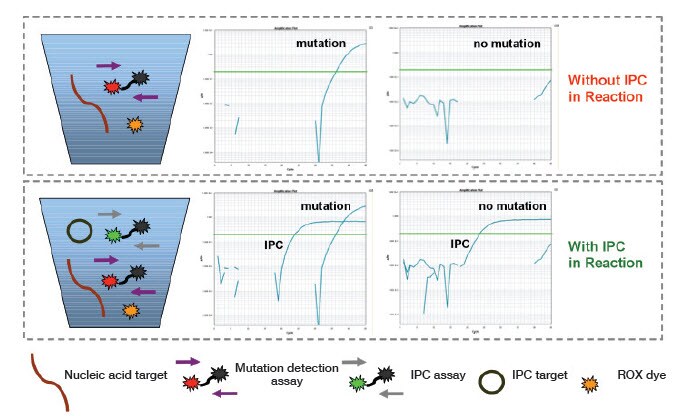
Check the Ct values of these samples with the gene reference assay. The Ct value determined for a gene reference assay on a sample should be approximately 18–28 for a 20 µL reaction, and 17–27 for a 10 µL reaction; these ranges correspond to a functional gDNA template input range of approximately 100 ng to 1 ng. If the Ct is higher than this, it indicates that the sample may be of poor quality (either degraded gDNA or inhibitors). You may have to use less DNA (10–50 ng; recommended input 20 ng). You might also want to consider using the IPC reagents in a duplex with your assay, to help identify false negatives.
Software Data Analysis
Make sure that paired samples have the same sample name when run with both the mutant and gene reference (or wild type) assay. Also, make sure the wild type sample run with the mutant assay is set as a negative control in the software.
Make sure the “Show % Mutation” Box is checked in the upper right-hand corner of the software. By default it is unchecked, so you must check it in order to see percent mutation.
The software accepts only *.txt or *.csv files from your exported data file. Make sure they are in one of these formats. Also, make sure you are following the correct naming conventions for the assays in the instrument software. The Mutation Detector™ Software first looks in the Assay Attribute file to get the pairing between mutant assay and corresponding gene reference assay. If the mutant allele assay does not exist in the Assay Attribute file, then it uses the assay names to pair the mutant allele assays with corresponding gene reference assays. The assay pairing information is used for mutation detection analysis, and to associate the assays with assay-specific attributes (for example, calibration ΔCt and detection ΔCt cutoff values).

You will need to edit the default Assay Attribute file to include your custom assay and the calibration ∆Ct value. Save the text file under a new name and then load the custom file you created with your new study. Make sure to follow the same format for your assay: Gene_###_mu or Gene_###_wt or Gene_rf. Custom assays using a mutant and wild type pairs are meant to be analyzed with a calibration ∆Ct value to determine percent mutation.
This is the value used in the ΔCt cutoff calculations. For any Ct values that are undetermined or greater than the maximum Ct cutoff value (set in the Current Study Settings), the maximum Ct cutoff value is shown and is used for the ΔCt cutoff calculations. The default Ct cutoff value is 37; you can set your own Ct cutoff value within a range of 30–40. Note: Ct cutoff values >37 may lead to false positive results in some cases.
For Research Use Only. Not for use in diagnostic procedures.