Search Thermo Fisher Scientific
StepOne™and StepOnePlus™ Real-Time PCR Systems Support – Getting Started
Find valuable information.
Optimize your experiments to get the best results. We’ve compiled a detailed knowledge base of the top tips and tricks to meet your research needs.
View the relevant questions below:
Calibration
| Recommended Maintenance Schedule | |
|---|---|
| Power on/off the computer controlling the instrument | Weekly |
| Check computer disk space. If necessary, archive or back up your experiment files and instrument settings | Weekly |
| Background calibration | Every month |
| Run disk cleanup and disk defragmentation | Every month |
| Perform an instrument self test | Every month |
| Pure dye calibrations | Every 18 months |
| Spatial calibration | Every 18 months |
| RNaseP instrument verification | After the instrument has been moved, or as needed to verify instrument performance |
The RNaseP verification plate contains template, master mix, and a TaqMan® assay for RNaseP. It is used to verify that the instrument is performing to specifications. If you have reason to suspect there is something wrong with the instrument, or if you want to rule out a chemistry issue, the RNaseP plate is a good way to test the system. It is also recommended to use after the instrument has been moved to a new location. The RNaseP verification plate is a single-use plate.
The calibration plates can be stored and reused three times for up to one year after you receive them, so make sure to return them to their original packaging and return them to –20°C storage until the next use. If needed, you can make your own background plate using deionized water. Please follow the directions in the StepOne™ and StepOnePlus™ Maintenance and Administration Guide (Appendix C) for more details.
Operation
The StepOne™ and StepOnePlus™ Real-Time PCR Systems have a Fast block, which supports reaction volumes of 10–30 µL.
Follow these instructions for proper loading of tubes, tube strips, and plates.
- Put the tubes or tube strips in the tray with the adapter on the 96-well support base.
- If using a plate, put the plate directly on the 96-well support base.
- Pipet your reactions into the tubes/tube strips or the wells on the plate.
- Seal the tube/tube strips with optical flat caps.
- If using a plate, seal the plate with either optical adhesive film or flat caps.
For optimal performance on the StepOnePlus™ instrument with partial loads:
- Load at least 16 tubes and arrange them in adjacent columns of 8 tubes, using rows A through H, or adjacent rows of 8 tubes, using columns 3 through 10.
For optimal performance on the StepOne™ instrument, load at least 4 tubes in the sample block.
Please refer to this selection table for compatible plates, tubes, and films.
The StepOnePlus™ Real-Time PCR System contains six independently thermally regulated VeriFlex™ blocks, which can help you optimize your thermal cycling conditions. You can set a different temperature for one or more zones, or you can set one temperature for all zones in the sample blocks. [Note: The difference in temperature between adjacent zones can range from 0 to 5°C, in 0.1°C increments].
- Choose ‘New Experiment’ → ‘Advanced Setup’ and choose the Experimental Properties as normal.
- Under ‘Plate Setup’ → ‘Assign Targets and Samples’
- On the right-hand side under ‘View Plate Layout’, check the box to ‘Enable VeriFlex™ Block’, and click ‘Yes’ to the box. You should now see the plate separated into 6 zones by red lines.

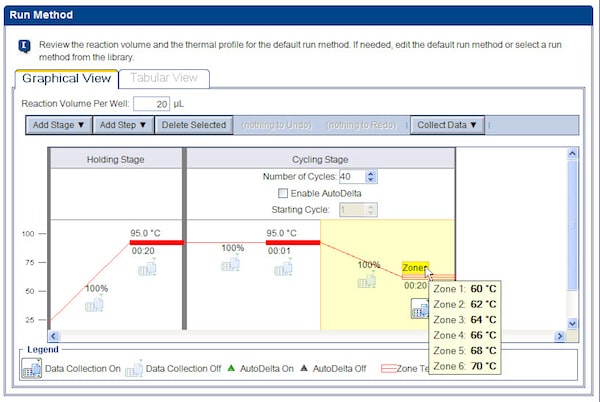
- Click on ‘Run Method’ in the left-hand column. On the Graphical View, click on the temperature that you want to change. A new window will pop up. Choose ‘Set different temperatures for one or more zones’. Adjust the temperature as needed for each zone.

- After you click ‘OK’ you will see the ‘Zones’ represented on the thermal profile. If you hover over the word ‘Zones’, you can see the temperatures you set.
In a touchdown PCR experiment, you will change either the temperature or the time of a particular PCR step with every cycle. Most commonly, the annealing temperature is adjusted throughout the experiment, such that the specificity is higher in the early cycles and the efficiency in the later cycles. In this example, we will set the method to do the following:
40 cycles  |
|
- Go to File → New Experiment → Advanced Setup. Fill out the relevant options as you normally would.
- Go to the Run Method under the ‘Setup’ section and you should see the Graphical View of your thermal profile. Check the box next to ‘Enable AutoDelta’. You should see some grey triangles appear next to the Temperature and Time at every step in the Cycling Stage. (Note: If you want to start the changes at a later cycle, set this here under ‘Starting Cycles’.)

- A new window called ‘AutoDelta Settings’ will open up. Select the appropriate options. In this example we are decreasing the temperature by 0.4°C per cycle, so choose (“-“) and (0.40). Click ‘Save Setting’. You will then see a green triangle show up next to the parameter you changed, in this case next to the 72°C step. Your new method has now been applied.

MicroAmp 96-Well Tray for VeriFlex Blocks (Cat. No. 4379983) is needed to load 1 or 2 MicroAmp Optical 8-Tube Strips with Attached Optical Caps on the ProFlex, SimpliAmp, MiniAmp, MiniAmp Plus, and Veriti Thermal Cycler 96-well blocks, and QuantStudio 3 and QuantStudio 5 blocks.
The MicroAmp Optical 8-Tube Strip with Attached Optical Caps is compatible with the following instruments:
- ProFlex 3 x 32-well PCR System
- ProFlex 96-well PCR System
- Veriti 96-well Thermal Cycler
- SimpliAmp Thermal Cycler
- MiniAmp Thermal Cycler
- MiniAmp Plus Thermal Cycler
- 2720 Thermal Cycler
- GeneAmp PCR System 9700, 96-well
- 7000 PCR System
- 7300 Real-Time PCR System
- 7500 Real-Time PCR system
- QuantStudio Real-Time PCR Systems
- ViiA7 Real-Time PCR System, 96-well
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
The MicroAmp 8-Tube Strip with Attached Domed Caps is compatible with the following instruments:
- ProFlex 3 x 32-well PCR System
- ProFlex 96-well
- Veriti 96-well Thermal Cycler
- SimpliAmp Thermal Cycler
- MiniAmp Thermal Cycler
- MiniAmp Plus Thermal Cycler
- 2720 Thermal Cycler
- GeneAmp PCR System 9700, 96-well
MicroAmp 96-Well Tray for VeriFlex Blocks (Cat. No. 4379983) is needed to load 1 or 2 MicroAmp Optical 8-Tube Strips with Attached Domed Caps on the ProFlex, SimpliAmp, MiniAmp, MiniAmp Plus, and Veriti Thermal Cycler 96-well blocks.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
Please refer to the table on Page 2 of this flyer for compatibility information.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
Please refer to the table on Page 2 of this flyer for compatibility information.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
Data Analysis
Run files will be saved to both the instrument and the connected computer. On the connected computer, files will be saved to the default data folder, unless you change it.
To find or change the default folder, go to Tools → Preferences → Defaults. Here you will see a Data Folder and an Import Folder. The default location is shown. If you want files to be saved to (or open from) a different location, click ‘Browse’ and choose the new folder.
For genotyping data, we recommend TaqMan® Genotyper Software. For relative quantitation, we recommend ExpressionSuite™ Software.
For Research Use Only. Not for use in diagnostic procedures.