Search
Automate Cells-to-CT™ Kits for High Throughput Expression Analysis
The Cells-to-CT™ Kits for TaqMan® and SYBR® Green-based real-time PCR take you from cells in culture to real-time RT-PCR analysis without RNA purification. The Cells-to-CT system uses a proprietary method for lysing cultured cells while simultaneously removing genomic DNA and preserving RNA integrity, saving significant time and effort over traditional methods requiring RNA purification. Here, Applied Biosystems scientists demonstrate the ease with which the Cells-to-CT methodology can be automated for high throughput gene expression analysis.
Simple Procedure Amenable to Automation
The Cells-to-CT™ Kits for TaqMan® and SYBR® Green-based real-time PCR make it possible to perform gene expression analysis using cultured cells without purifying RNA. Cells can be processed directly in 96- or 384-well culture vessels using a 2-step, 7 minute procedure that does not require the heating, centrifugation, or vacuum filtration procedures that are typically required for RNA isolation (Figure 1). The simplicity of the Cells-to-CT method makes it extremely amenable to use with even the most basic automated liquid handling systems. Automation reduces labor and plastic consumption, and minimizes the opportunity for cross-contamination, sample loss, and other human errors. As a result, automating the Cells-to-CT protocol provides researchers with an easier, more reproducible gene expression analysis workflow suitable for high throughput projects. Using commonly available robotic systems, the Cells-to-CT Kits can be automated to prepare 384 unique cell samples for real-time RT-PCR in less than 10 minutes—less than 1.5 seconds per sample!
Sample Mixing is Not Required
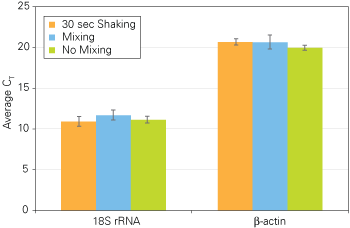
The protocols for manually processing cells with Cells-to-CT Kits recommend mixing samples after the addition of both the the Lysis and Stop Solutions. To facilitate automation of the procedure, we investigated whether these sample mixing steps are required for reproducible, high quality real-time PCR data using the TaqMan Gene Expression Cells-to-CT Kit. After the addition of Lysis and Stop Solutions, samples were either mixed by pipetting 5 times (the standard recommended protocol), shaken for 30 seconds, or not mixed. Very similar CT values and levels of reproducibility were obtained across these different sample mixing conditions (Figure 2), indicating that, with small samples of HeLa cells, the mixing step can be omitted from the Cells-to-CT sample preparation procedure. (Note: this data was generated using 4,000 HeLa cells, which are are relatively easy to lyse. Results may be different with other cell lines and different cell numbers. The no-mix protocol should be evaluated with the cell lines and quantities in the experimental system before conducting larger experiments). Eliminating these mixing steps not only makes the procedure even simpler and more rapid, but it has the added benefit of dramatically reducing the number of pipette tips needed for the procedure. The result is a faster, more economical procedure with fewer steps.
Reproducible Results
Automated on a Biomek® FX Workstation, the Cells-to-CT no-mix procedure results in gene expression data with equivalent sensitivity and reproducibility to the standard manual Cells-to-CT methodology, while reducing labor and minimizing cross-contamination. Figure 3 demonstrates the wide dynamic range, high sensitivity, and excellent reproducibility achieved with the automated procedure. Figure 4 demonstrates that cross contamination is minimized with the automated protocol. The resulting lysates are compatible with the full range of optimized Applied Biosystems TaqMan and SYBR Green PCR master mixes.
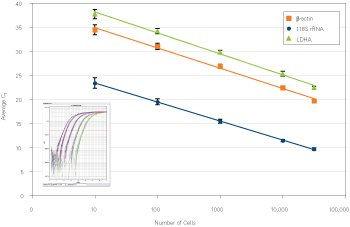
Figure 3. Automating the Cells-to-CT™ Procedure Results in Sensitive, Reproducible Expression Data. The average CT value of 16 biological samples per MCF-7 cell concentration on a 96-well flat-bottomed tissue culture plate was determined for ß-actin (orange squares), 18S (blue circles), and LDHA (green triangles) using the TaqMan® Cells-to-CT Kit and TaqMan Gene Expression Assays.
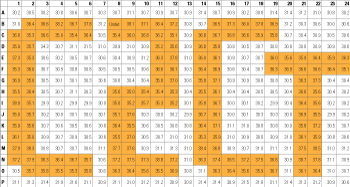
Figure 4. Cross Contamination is Minimized with the Automated Cells-to-CT™ Procedure. Real-time PCR for HPRT1 was carried out on 224 biological replicates (2,500 HeLa cells per well) interspersed with cell-free blanks (160 wells, arranged to spell out “C2CT”) in a 384-well flat-bottomed culture plate. The CT values for blank wells were >35 in all cases (orange wells) while the average CT value for the samples was 30.7 ± 0.5 showing that there was no cross-contamination between wells. ΔCT (HPRT1–ß-actin) of wells containing cells was 5.4 ± 0.4—this low error indicates that differences in CT values across the plate are due to differences in cell number rather than to real-time PCR variation.
Conclusions
Automating the Cells-to-CT procedure results in a more efficient usage of time, labor, and plastics, yet maintains the highest quality gene expression data with equivalence to the standard methodology.
Scientific Contributors:
Ronald V. Abruzzese, Richard Fekete • Applied Biosystems
