Search
Step 2. Quantify functional DNA
Because DNA from FFPE tissues is heavily fragmented and chemically modified, only a small fraction is functional for genotyping. Determine how much is functional using TaqMan® RNase P Detection Reagents.
In FFPE tissue samples, nucleic acids become trapped and modified through protein-protein and protein-nucleic acid cross-links, and other chemical modifications. Furthermore, FFPE DNA is often fragmented to a range of sizes. It can have peak fragment sizes ranging from ~180 base pairs (bp) to sizes that are slightly smaller than those observed with frozen tissue (peak fragment size ~3000 bp).
Fragmentation of DNA can be easily assessed on a gel, but treatment with formaldehyde causes extensive modification of biomolecules that are not apparent when looking at size alone. These modifications affect the ability of DNA to function as an effective template, sometimes to the point that only a fraction of the available DNA obtained from FFPE samples is functional, i.e. can provide a signal using a polymerase-based assay such as PCR. To quantify the level of functional DNA, we recommend targeting the single-copy gene RNase P in a Real-time PCR assay. If the percentage of functional DNA is too low, even this solution will not be adequate (possibly due to effects from the “junk” DNA), so quantifying the DNA using A260 measurements is also necessary. If the amount of DNA determined by the two methods varies by more than 20-fold (i.e., less than 5% of the DNA is determined to be PCR-functional), the PCR analysis may be compromised.
>> Recommendation: Perform DNA quantification using A260 measurements along with TaqMan® RNase P Detection Reagents (FAM™) to determine the percentage of functional template in a DNA sample. The probability of successful PCR analysis is greatly reduced when the amount of the functional template in the FFPE DNA sample is less than 5%.
How to Quantify Functional DNA
Prepare a standard curve using the DNA template standards provided with the TaqMan® DNA Template Reagents and the RNase P gene primers and probe provided with the TaqMan® RNase P Detection Reagents (FAM™). Prepare the PCR Mix using 2X TaqMan® Universal PCR Master Mix, No AmpErase® UNG, and 20X RNase P Primer and TaqMan Probe (FAM™ Dye) mix. It is important to perform at least 3 replicates of each standard or sample and to include a no-template control. Combine 10 ng of FFPE DNA and PCR mix in either MicroAmp™ 96-Well Optical Reaction Plates or MicroAmp™ 384-Well Clear Optical Reaction Plates. Run the plated reactions on an ABI Prism® Sequence Detection System or 7900 HT Fast Real-Time PCR System using the following thermal cycling conditions in Standard mode: hold for 10 min at 95°C, then 40 cycles with denaturation for 15 sec at 92°C and anneal/extend for 1 min at 60°C. Finally, use the Sequence Detection System or PCR System software to generate a standard curve to quantify the amount of functional template and, subsequently, calculate the percentage of functional template in each FFPE DNA sample.
Case Study:
DNA Quantification
Both A260 measurements and TaqMan® RNase P Detection Reagents (FAM™) were used to assess the amount of functional DNA in each FFPE DNA sample (as described above in “How to Quantify Functional DNA”).
The average yield of DNA/mg tissue, as determined by A260 measurements, was 1.26 ± 1.47 µg. This demonstrates the wide variability of yields from FFPE tissue.
10 ng (as quantified by A260 measurements) of each sample was then assessed with the RNase P assay. The average percentage of functional template was 10.07 ± 7.08%, indicating that A260 values are not a reliable gauge of functional templates for FFPE samples. RNase P detection qPCR data (not shown) demonstrated that the RecoverAll™ Total Nucleic Acid Isolation Kit can provide a reproducible source of DNA with ± 0.5 Ct and ± 2% functional template difference between replicate isolations from any one tissue.
All assays in this series of experiments were successfully performed with 1 ng of DNA input (as determined by RNase P detection), which is considered the lower limit of sample quantity required for TaqMan® SNP Genotyping Assays. Since the SNP assay requires 1 ng in 2.25 µL, samples that contained a low percentage of functional template were concentrated in a centrifugal evaporator.
It is critical for the success of SNP genotyping that each sample contains the same concentration of functional DNA template. When comparing human FFPE DNA quantification methods (A260 vs. RNase P), the SNP assays based on RNase P quantification produced tighter clustering of genotype groups on allelic discrimination plots after 40 cycles of amplification (Figure 1 below, compare Panels A and B).
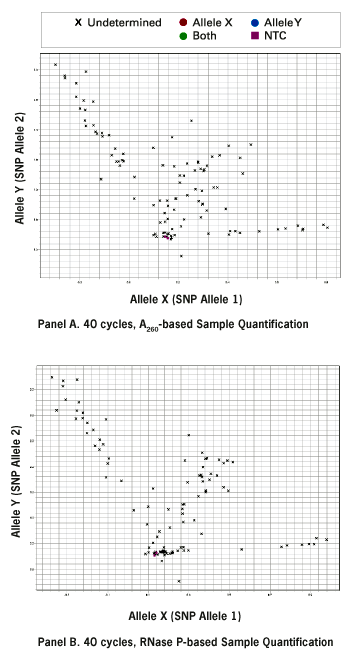
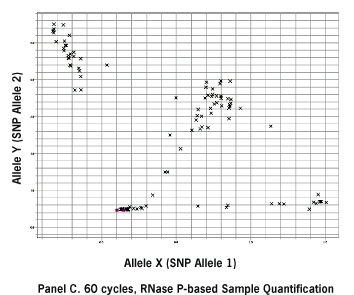
Figure 1. Allelic Discrimination Plots Comparing FFPE DNA Quantification Methods and Number of Amplification Cycles. 109 human FFPE DNA samples (1 ng each) were amplified for 40 cycles (Panels A, B) or 60 cycles (Panel C) and AutoCalled using Sequence Detection Systems 2.2.1 Software. (A) Sample input based on UV absorbance (A260) quantification. (B), (C) Sample quantification by RNase P detection.