Search
7500 and 7500 Fast Real-Time PCR Systems Support - Getting Started
Find valuable information
Optimize your experiments to get the best results. We’ve compiled a detailed knowledgebase of the top tips and tricks to meet your research needs.
View the relevant questions below:
Having problems with your experiment? Visit our
| Recommended Maintenance Schedule | |
|---|---|
| Power on/off the computer controlling the instrument | Weekly |
| Check computer disk space. If necessary, archive or back up your experiment files and instrument settings | Weekly |
| Background calibration | Every month |
| Run disk cleanup and disk defragmentation | Every month |
| Perform an instrument self test | Every month |
| Pure dye calibrations | Every 6 months |
| ROI calibration | Every 6 months |
| Optical calibration | Every 6 months |
| RNaseP instrument verification | After the instrument has been moved, or as needed to verify instrument performance |
The calibration plates can be stored and reused three times for up to 12 months after you first open them, so make sure to return them to their original packaging and return them to –20°C storage until the next use. If needed, you can make your own background plate using deionized water. Please follow the directions in the 7500 and 7500 Fast Real-Time PCR System Maintenance Guide (Appendix C) for more details.
The RNaseP verification plate contains template, master mix, and a TaqMan® assay for RNaseP. It is used to verify that the instrument is performing to specifications. If you have reason to suspect there is something wrong with the instrument, if the instrument has been moved, or if you want to rule out a chemistry issue, the RNaseP plate is a good way to test the system. The RNaseP verification plate is a single-use plate.
Follow the directions in the 7500 and 7500 Fast Real-Time PCR System Maintenance Guide, Chapter 6, to decontaminate the sample block. You can also watch this helpful video demonstration.
| Instrument Specifications | |
|---|---|
Block Options | 7500: 96-well (standard) |
| Sensitivity | Down to 1 copy |
| Dynamic Range | 9 logs of linear dynamic range |
| Calibrated Dyes | FAM™, SYBR®, VIC®, ROX™, NED™, TAMRA™ dyes (Cy®3, Cy®5, and Texas Red® dyes – 7500F only) |
| Detection Method | SYBR® dye, primer-probe detection |
| Resolution | Detect changes as little as 1.5-fold |
| Reaction Volume Range | 20–100 µL (7500) |
| Reaction Speed | Fast or standard (7500 has standard mode only) |
| Optics | Tungsten-halogen lamp, 5 excitation filters, 5 emission filters, CCD Camera |
| Temperature Range | 4–99.9°C |
| Run Time | <2 hr (standard mode) |
| Regulatory Statement | For Research Use Only. Not for use in diagnostic procedures. |
| Temperature Accuracy | ±0.25°C (between 35°C and 95°C, after 3 min) |
| Temperature Uniformity | ±0.5°C (after 30 sec) |
| Thermal Cycling System | Peltier-based system |
| Available Applications | Gene expression, genotyping, copy number variation, HRM, protein thermal shift, protein detection, mutation detection, miRNA, presence/absence |
| Dimensions | 34 cm (W) x 45 cm (D) x 49 cm (H) |
| Weight | 34 kg (75 lb) |
| Remote Monitoring | No |
| On-Board Memory | No |
| Setup Configurations | PC-controlled only |
The Applied Biosystems® 7500 and 7500 Fast Real-Time PCR Systems use the following dye sets for calibration: Cy®3, Cy®5, FAM™, JOE™, NED™, ROX™, SYBR® Green, TAMRA™, Texas Red®, and VIC® dyes. The following figure shows the emission spectrum for each dye, and the filters and wavelengths at which each dye is read. Custom dyes that are read between 520 and 650 nm can also be used, although you will have to calibrate the system first for any new dye.

The 7500 and 7500 Fast Real-Time PCR Systems can be used to run assays designed with custom dyes (dyes not manufactured by Life Technologies). Custom dyes must fluoresce within the 520–650 nm spectral range measured by the 7500 or 7500 Fast instrument. To use a custom dye, you must first determine what the right concentration of the dye is. You will need to order an oligo with a 5’ custom dye but no quencher. Make up a plate with different concentrations of this oligo (~25–3,200 nM) and use the ROI Inspector to assess fluorescence. Choose the concentration that displays the brightest possible signal without saturation in all filters. Once you have found the correct concentration, create a full plate of custom dye at this concentration and perform the custom dye calibration. See Appendix B in the 7500 and 7500 Fast Real-Time PCR System Maintenance Guide for full details.

Using the instrument computer when a run is in progress is not recommended, as this poses a risk of corrupting the data.
The following volumes are supported for each instrument block:
- 7500: 20–100 µL reactions
- 7500 Fast: 10–30 µL reactions
No. The software will use the last data collection step in the cycling stage for all amplification plots and Ct analysis. So even if you were to set two separate steps with “Data Collection On”, you would only be able to view and analysis data from the latter step.
Run files will be saved to a default folder on the connected computer, unless you change it. To find or change the default folder (in SDS v2.0.1 or later), go to Tools → Preferences → Defaults. Here you will see a Data Folder and an Import Folder. The default location is shown. If you want files to be saved to (or open from) a different location, click ‘Browse’ and choose the new folder.
In a touchdown PCR experiment, you will either change the temperature or the time of a particular PCR step with every cycle. Most commonly, the annealing temperature is adjusted throughout the experiment, such that the specificity is increased in the early cycles and the efficiency in the later cycles.
In this example, we will set the method to do the following:

- Go to File → New Experiment → Advanced Setup. Fill out the relevant options as you normally would.
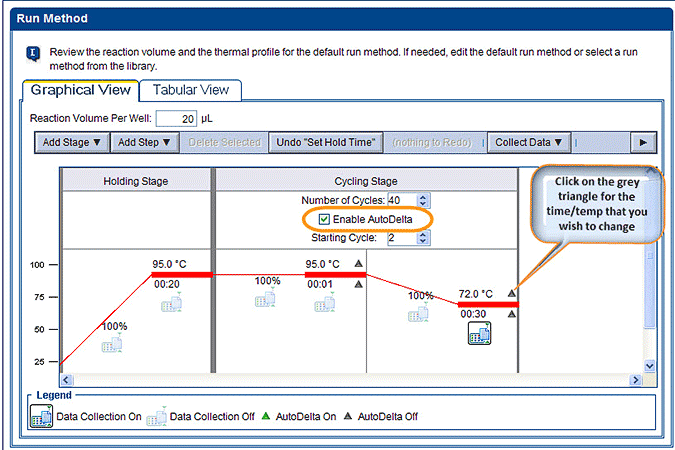
- Go to the Run Method under the ‘Setup’ section and you should see the Graphical View of your thermal profile. Check the box next to ‘Enable AutoDelta’. You should see some grey triangles appear next to the Temperature and Time at every step in the Cycling Stage. (Note: If you want to start the changes at a later cycle, set this here under ‘Starting Cycles’.)
- A new window called ‘AutoDelta Settings’ will open up. Select the appropriate options. In this example we are decreasing the temperature by 0.4°C per cycle, so choose (“-“) and (0.40). Click ‘Save Setting’. You will then see a green triangle show up next to the parameter you changed, in this case next to the 72°C step. Your new method has now been applied.

Yes. If you have the newer version of the software (v2.0.1 or later), which creates *.eds files, your data will be directly compatible with our Protein Thermal Shift™ Software. If you have the older software (v 1.x), you will have to program the software differently (see below) and analyze the results independently. For more details on the analysis, you can refer to this paper: “The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability.” Nat Protoc 2007;2(9):2212-21.
For an experiment using SDS v1.x, follow the directions below:
- Create a new experiment, choosing Absolute Quantification assay type. Click ‘Next’.
- Go to ‘Select Detectors’. If there is no predefined detector for this application, create a new one. Make sure the Reporter Dye is ROX, and Quencher Dye is none.
- Assign the detector to the plate. Select none for the Passive Reference.

- Next go to the Instrument tab and change the thermal profile as follows:

- Go to the Auto Increment:

MicroAmp 96-Well Tray for VeriFlex Blocks (Cat. No. 4379983) is needed to load 1 or 2 MicroAmp Optical 8-Tube Strips with Attached Optical Caps on the ProFlex, SimpliAmp, MiniAmp, MiniAmp Plus, and Veriti Thermal Cycler 96-well blocks, and QuantStudio 3 and QuantStudio 5 blocks.
The MicroAmp Optical 8-Tube Strip with Attached Optical Caps is compatible with the following instruments:
- ProFlex 3 x 32-well PCR System
- ProFlex 96-well PCR System
- Veriti 96-well Thermal Cycler
- SimpliAmp Thermal Cycler
- MiniAmp Thermal Cycler
- MiniAmp Plus Thermal Cycler
- 2720 Thermal Cycler
- GeneAmp PCR System 9700, 96-well
- 7000 PCR System
- 7300 Real-Time PCR System
- 7500 Real-Time PCR system
- QuantStudio Real-Time PCR Systems
- ViiA7 Real-Time PCR System, 96-well
p>Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
The MicroAmp 8-Tube Strip with Attached Domed Caps is compatible with the following instruments:
- ProFlex 3 x 32-well PCR System
- ProFlex 96-well
- Veriti 96-well Thermal Cycler
- SimpliAmp Thermal Cycler
- MiniAmp Thermal Cycler
- MiniAmp Plus Thermal Cycler
- 2720 Thermal Cycler
- GeneAmp PCR System 9700, 96-well
MicroAmp 96-Well Tray for VeriFlex Blocks (Cat. No. 4379983) is needed to load 1 or 2 MicroAmp Optical 8-Tube Strips with Attached Domed Caps on the ProFlex, SimpliAmp, MiniAmp, MiniAmp Plus, and Veriti Thermal Cycler 96-well blocks.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
Please refer to the table on Page 2 of this flyer for compatibility information.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
Please refer to the table on Page 2 of this flyer for compatibility information.
Yes. The tubes are DNase/RNase/Human DNA free, as they are manufactured in a Class 100K ISO certified clean room UK production facility.
These new strips have new features including a graduated 20 µL measuring mark for visual verification and dual end tabs for better labeling and handling. In addition, the individual caps help prevent cross contamination and reduce sample evaporation.
- Go to ‘View Well Table’ from the Amp Plot screen.
- Click on the ‘Group By’ button and choose ‘Well Position’ (Column). Then highlight the plate and print the report.
You can also Export Data, and the column format will be retained.
For genotyping data, we recommend TaqMan® Genotyper Software. For relative quantitation, we recommend ExpressionSuite™ Software. (If you have SDS v1.x, you would have to use DataAssist™ software for relative quantitation data.)
For Research Use Only. Not for use in diagnostic procedures.