Search
Synthesize High Yields of Biotinylated aRNA
Ambion now offers an improved MessageAmp™ II Kit with biotin-11-UTP. MessageAmp II-Biotin Enhanced provides a complete single round amplification kit for biotin array analysis. The biotin modified UTP and ratio of labeled to unlabeled UTP were selected based on superior performance and aRNA yield. The biotinylated nucleotide is supplied as a pre-mixed NTP at an optimized ratio, providing convenient,
consistent labeling.
- Highest amplification in a single round—up to 10,000-fold for most samples
- Enough aRNA for Affymetrix® GeneChip® analysis from just 50 ng total RNA
- Optimized NTP mix containing biotinylated UTP reduces set-up time and maximizes labeling
- Reduce costs by 45%
T7 RNA Polymerase Linear Amplification is the Gold Standard
Microarrays measure the expression levels of thousands of genes. One limitation of this technology, however, is the need for large amounts of labeled target RNA for hybridization to the microarray. This limitation is overcome by using a T7 linear amplification protocol commonly referred to as the “Eberwine Method.” This method most accurately maintains the initial mRNA expression profile and has been the gold standard for microarray target preparation [1]. Biotin, incorporated as biotinylated nucleotide triphosphates during the in vitro transcription (IVT) step, is the preferred label for most commercial microarray platforms. The highly specific and efficient binding of streptavidin-dye conjugates (phycoerythrin, Cy™ dyes) to biotinylated RNA/oligonucleotide probe hybrids is the basis of highly sensitive fluorescence detection. Ambion has incorporated biotin labeling reagents into the MessageAmp II Kit to provide a complete microarray aRNA preparation kit, the
MessageAmp II-Biotin Enhanced Kit.
Higher Yields of Labeled aRNA; Only 1 Round of Amplification
Both biotin incorporation and further optimization of the IVT reaction has resulted in increased sensitivity of microarray transcript detection. The increased sensitivity allows the use of less input RNA-50 ng of input RNA in a single round of amplification provides enough aRNA for array analysis. Amplified RNA yields derived from nine different tissue and cell sources and four different input amounts are shown in Figure 1. While length of the aRNA synthesized will largely depend on RNA sample quality, the cDNA and IVT reaction conditions in this kit produce longer biotinylated aRNA compared to the market leader. Figure 2 shows comparable Agilent® bioanalyzer scan results from labeled aRNA produced with MessageAmp II-Biotin Enhanced Kit and unlabeled aRNA produced with MessageAmp II Kit. The resulting improvements also give the MessageAmp II-Biotin Enhanced Kit a higher number of Present calls than the market leader (Figure 3).
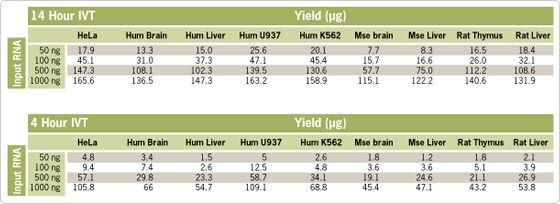
Figure 1. aRNA Yield from Nine Different Sources. Four different amounts of total RNA from nine different sample types were amplified using the MessageAmp™ II-Biotin Enhanced Kit. Average aRNA yields from triplicate reactions are shown for 4 or 14 hr in vitro transcription (IVT) reactions. These data are useful for determining both the amount of total RNA needed to obtain enough labeled aRNA for array hybridization (typically ~10 µg) and the optimal length of IVT incubation. Note that there is a ~3-fold difference in aRNA yield between some samples. With most RNA sources, 50–100 ng of input total RNA amplified with the MessageAmp II-Biotin Enhanced Kit using a 14 hr IVT incubation will yield enough labeled aRNA for microarray hybridization.
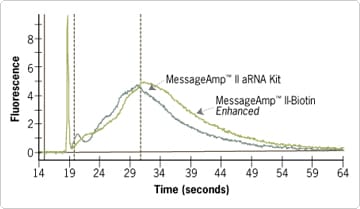
Figure 2. Electropherogram of Biotin-labeled vs. Unlabeled aRNA. Identical HeLa cell total RNA samples (1000 ng) were amplified with either the MessageAmp™ II-Biotin Enhanced Kit to produce biotin-labeled aRNA or the MessageAmp II Kit to produce unlabeled aRNA. The IVT reactions were carried out for 4 hr. Equivalent mass amounts from each reaction were analyzed on an Agilent® bioanalyzer. The shift in aRNA size from the MessageAmp II-Biotin Enhanced sample results from the incorporation of biotin into the aRNA.
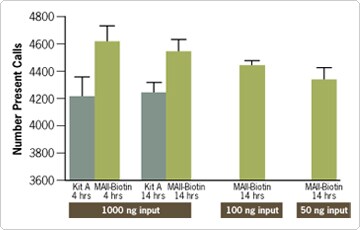
Figure 3. Number of Present Calls Detected on Affymetrix® Human Genome Focus Arrays. The indicated amounts of total RNA from HeLa cells was amplified in triplicate using either the MessageAmp™ II-Biotin Enhanced Kit (MA II-Biotin) or another manufacturer’s aRNA labeling kit (Kit A). Two IVT incubation times (4 and 14 hrs) were tested with 1000 ng of total RNA input. Amplification of as little as 50 ng total RNA with the MessageAmp II-Biotin Enhanced Kit produced a comparable number of Present calls as 1000 ng input RNA that was amplified with Kit A. Furthermore, the amplification reaction can be completed in one day.
Hybridization of the biotinylated aRNA produced from the MessageAmp II-Biotin Enhanced Kit to Affymetrix GeneChip Arrays was used to assess the sensitivity and reproducibility of the method and reagents. Increased sensitivity can be measured several ways. One indication of sensitivity is the number of Present calls calculated by the GeneChip Operating Software (Figure 3). Additional analysis of the distribution of signal intensities indicates that the new MessageAmp II-Biotin Enhanced Kit produces a higher mean value. This means that a there was a higher signal intensity across the array (Figure 4). This results in more reproducible array results and increased sensitivity of lower expressed transcripts.
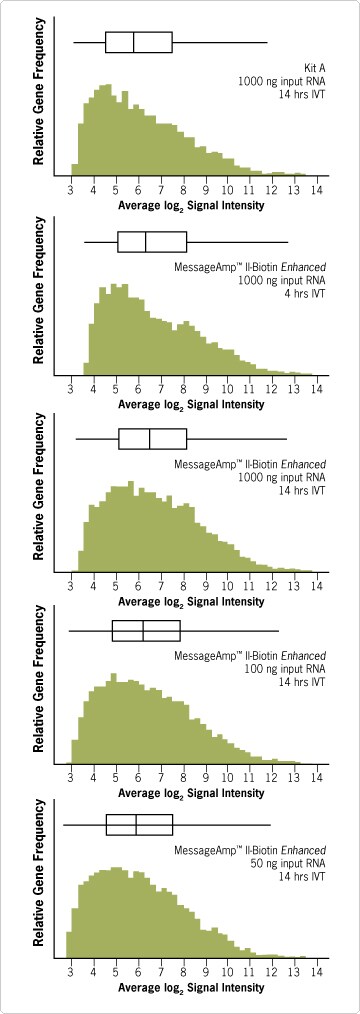
Figure 4. Higher Signal Intensity Distribution on Affymetrix® Human Genome Focus Arrays. Distributions of log2 signal intensity values were plotted for the MessageAmp™ II-Biotin Enhanced Kit and another manufacturer’s amplification kit (Kit A). The median values for each of the MessageAmp II-Biotin Enhanced plots were similar for total RNA inputs ranging from 50 to 1000 ng. In contrast, signal intensities produced by Kit A at the 1000 ng total RNA input level was markedly reduced. The box and whiskers plot above each histogram indicate the median value (vertical center line), interquartile range (left and right sides of box), and the 10% and 90% quantiles (ends of line). Each plotted distribution represents the average log2 signal intensity of triplicate target preparations and hybridizations. IVT reactions were carried out for 4 or 14 hrs as indicated. The data were normalized within each group using Robust Multichip Average (RMA) (See sidebar, Data Analysis: Robust Multichip Average (RMA) and Affymetrix Algorithms).
Biotin Incorporation
Biotin-11-UTP serves as the label in the MessageAmp II-Biotin Enhanced Kit. Only a single biotinylated NTP is used because studies at Ambion and elsewhere [2] showed that in previous dual labeling experiments using biotin-UTP and biotin-CTP, the signal contributed by biotin-CTP was minimal, and relative signal intensity between genes was not altered with the use of a single biotin UTP label.
While various linker arm sizes (e.g., Biotin-11-UTP, Biotin-16-UTP) did not affect modified UTP incorporation into aRNA during IVT, the longer linker arms slightly impeded subsequent purification of the aRNA. Biotin-11-UTP provided the highest yields of aRNA.
The ratio of biotin UTP to unlabeled UTP was optimized to provide the highest yield of aRNA and the highest signal:noise on Affymetrix GeneChip Arrays. A fairly wide range of concentrations of Biotin-UTP (25–60%) can be used for acceptable detection, whereas at lower concentrations the signal drops dramatically, and at higher concentrations both the noise increases and the aRNA yield decreases.
Ambion realizes that microarray users may prefer to continue using their own biotin-labeling method, or to experiment with different labeling conditions. To accommodate this we offer two Biotin-UTP nucleotides (Biotin-11-UTP and Biotin-16-UTP) separately, which can be used with the standard MessageAmp II Kit.
While various linker arm sizes (e.g., Biotin-11-UTP, Biotin-16-UTP) did not affect modified UTP incorporation into aRNA during IVT, the longer linker arms slightly impeded subsequent purification of the aRNA. Biotin-11-UTP provided the highest yields of aRNA.
The ratio of biotin UTP to unlabeled UTP was optimized to provide the highest yield of aRNA and the highest signal:noise on Affymetrix GeneChip Arrays. A fairly wide range of concentrations of Biotin-UTP (25–60%) can be used for acceptable detection, whereas at lower concentrations the signal drops dramatically, and at higher concentrations both the noise increases and the aRNA yield decreases.
Ambion realizes that microarray users may prefer to continue using their own biotin-labeling method, or to experiment with different labeling conditions. To accommodate this we offer two Biotin-UTP nucleotides (Biotin-11-UTP and Biotin-16-UTP) separately, which can be used with the standard MessageAmp II Kit.
MessageAmp II-Biotin Enhanced Kit Format
The MessageAmp II-Biotin Enhanced Kit provides enough reagents for 20 single round amplification reactions. It can be used in combination with the
MessageAmp II Kit for small samples that require 2 rounds of amplification (MessageAmp II for the first round and MessageAmp II-Biotin Enhanced in the second round for labeling). Biotin-UTP is provided premixed with the other rNTPS for convenience and consistent, optimized labeling.
Scientific Contributors
Robert Setterquist, Mike Wilson, Charles Johnson, Shika Agarwal, Sharmili Moturi • Ambion, Inc.
Scientific Contributors
Robert Setterquist, Mike Wilson, Charles Johnson, Shika Agarwal, Sharmili Moturi • Ambion, Inc.
References1. Van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD, Eberwine JH (1990) Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc Natl Acad Sci USA 87(5):1663–7.
2. Dorris DR, Ramakrishnan R, Trakas D, Dudzik F, Belval R, Zhao C, Nguyen A, Domanus M, Mazumder A (2002) A highly reproducible, linear, and automated sample preparation method for DNA microarrays. Genome Res 12(6):976–84.
2. Dorris DR, Ramakrishnan R, Trakas D, Dudzik F, Belval R, Zhao C, Nguyen A, Domanus M, Mazumder A (2002) A highly reproducible, linear, and automated sample preparation method for DNA microarrays. Genome Res 12(6):976–84.
Data Analysis: Robust Multichip Average (RMA) and Affymetrix® AlgorithmsAll Affymetrix CEL files were processed using a combination of Robust Multichip Average (RMA) and Affymetrix MAS 5.0 algorithms. The background subtraction, expression summary, normalization, and log2 transformation of gene signals was carried out using the RMA method developed by Irizarry and colleagues [1] and Affymetrix MAS 5.0 with a scaling factor of 500. Irizarry et al. demonstrated that RMA has better precision for lower expression values and provides a greater than 5-fold reduction of the within-replicate variance as compared to MAS 5.0. RMA also provided more consistent estimates of fold change and provided higher specificity and sensitivity when using fold change analysis to detect differential expression. For this reason, RMA processed signal was used in all differential gene analysis. Present/Absent determination for each gene was calculated using the One-sided Wilcoxon’s Signed Rank test using the Affymetrix default settings. JMP® Statistical Analysis Software was used to build the distribution plots.
References
1. Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP (2003) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31(4):e15.
References
1. Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP (2003) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31(4):e15.