Search
LanthaScreen Tb Deconjugation Assay Reagents
Deconjugating enzymes (DUBs, Ubiquitin-like (UBL) cleaving enzymes) specifically cleave peptide or isopeptide bonds at the ubiquitin or UBL C-terminus. These proteases are believed to play both “housekeeping” functions, by maintaining pools of active ubiquitin, and regulatory functions, by rescuing specific target proteins from degradation by the proteasome via removal of ubiquitin from the target protein. The LanthaScreen Tb deconjugating assay tools provide a sensitive and general method for assaying deconjugating enzyme activity with excellent Z´-factor values.
The LanthaScreen Tb DUB Substrate, and the corresponding De-SUMO and De-NEDD8 Substrates, consist of an N-terminal yellow fluorescent protein (YFP) fusions to ubiquitin (or UBLs) with a short C-terminal extension containing an engineered cysteine residue. This engineered cysteine residue is labeled with a LanthaScreen terbium chelate to create the TR-FRET based deconjugation substrate. In the absence of a DUB, the LanthaScreen Tb DUB Substrate remains intact and exhibits a high degree of FRET. When the substrate is cleaved at the ubiquitin C-terminus by the enzyme, the FRET signal decreases (Figure 1). Similar assays have been developed for the UBL cleavage enzymes.
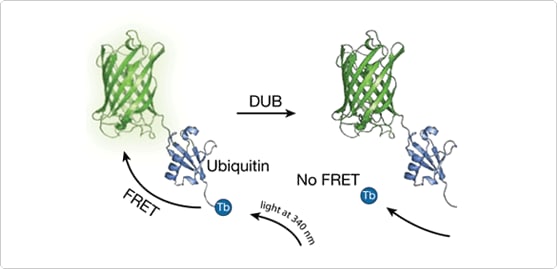 |
Figure 1. Mechanism of the LanthaScreen deconjugation assay. |
Due to the TR-FRET readout, the LanthaScreen deconjugating assays are ideally suited for screening compound libraries and examining the potency and selectivity of inhibitors (Figure 2) in either endpoint or kinetic assays (Figure 3).
 | | Figure 2. Influence of fluorescent compounds on the signal of ubiquitin–AMC and the LanthaScreen DUB Substrate. |
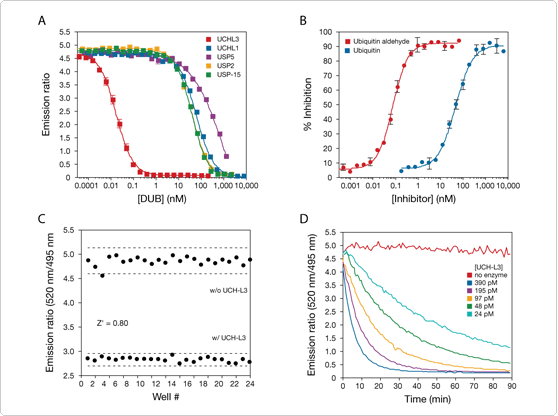
Figure 3. Endpoint and kinetic data readouts for the LanthaScreen DUB Substrate
Posters
For Research Use Only. Not for use in diagnostic procedures.