Search
RNase A Molecules are Used to Study the Structure of Amyloid-like Fibrils
Ambion’s RNaseAlert® Lab Test Kit is not only used to detect contaminating RNase activity in samples and reagents but is also useful in biological assays. In just 30 minutes, as little as 3.5 x 10-7 units (~0.5 pg) of RNase A activity can be detected.
RNaseAlert Tests for Assessing Protein Conformation:
Amyloid-like fibrils that form in vitro from soluble, native proteins are often studied as models for elucidating disease processes or for protein folding. Sambashivan and colleagues [1] recently published data that reconcile the idea that the native structure of functional proteins has the lowest free energy of all possible configurations with the observation that many normally soluble proteins can form amyloid fibrils that are also very stable (i.e., resistant to degradation and disassembly by chemicals or proteases). Modified RNase A molecules were used to show that proteins are not completely refolded to create fibrils, but instead, retain features of the native protein that have RNase activity measurable with RNaseAlert technology.
The Significance of Amyloid
Amyloid is a general term for protein deposits in tissues that share at least three characteristics: (1) extracellular, nonbranching fibrils, (2) affinity for Congo red, an aromatic dye, that results in changes in fluorescence intensity, and (3) cross-beta sheet quaternary structure that creates a characteristic X-ray diffraction pattern. It is still not known whether amyloid plaques are a cause or symptom of diseases like systemic amyloidosis; diabetes mellitus type 2; Alzheimer’s, Parkinson’s, or Huntington’s disease; and spongiform encephalopathies (prion diseases). Understanding the structure and formation of amyloid fibrils may advance disease prevention or treatment options, but fibrillar features (e.g., large size and variable length) have made the structure difficult to study. In a carefully designed study, Sambashivan and colleagues [1] successfully used modified RNase A molecules to examine protein conformation within fibrils.
Advantages of Using RNase A
Four characteristics of RNase A make it useful for studying amyloid-like fibril structure. First, each RNase A monomer contains four disulfide bonds which limit conformational changes. Second, RNase activity can be easily measured with a fluorescence-based assay (see box below, More about the RNaseAlert® Lab TestKit & QC System). Third, two essential catalytic histidine residues (H12 and H119) are located on separate domains (core domain and carboxy-terminal b-strand) that are linked by a hinge loop. Fourth, RNase A can be manipulated in vitro (freeze-dried from acetic acid) to form dimers, in which the carboxy-terminal b-strand interacts with the core domain of its partner [2].
Detecting RNase Activity in Amyloid-like Fibrils
The last two characteristics described above were exploited to create two inactive mutants (histidine to alanine substitutions: H12A and H119A) that each had amyloidogenic sequences (ten glutamine residues flanked by glycine residues: Q10) in place of the hinge loop. Each mutant was shown to form amyloid-like fibrils by Congo red and X-ray diffraction analyses, and the resulting fibrils did not have RNase activity. When mutants were mixed and allowed to form fibrils together, RNase activity was detectable. This indicated that the histidine in the nonmutated carboxy-terminal b-strand of the Q10-H12A mutant can interact with the histidine in the nonmutated core domain of the Q10-H119A mutant to create an RNase A active site.
The Proposed Model
More About the RNaseAlert® Lab Test Kit & QC System
Sambashivan et al. [1] used Ambion’s RNaseAlert® technology (patent pending) to assay biological activity in their structural biology research, a very specialized application. However, RNaseAlert is also useful in everyday experiments for fast, sensitive detection of RNase contamination that could interfere with RNA-based experiments (Figure 1). The RNaseAlert tests are sensitive to several RNases, including RNase A, RNase T1, RNase I, micrococcal nuclease, S1 nuclease, mung bean nuclease, and Benzonase®. 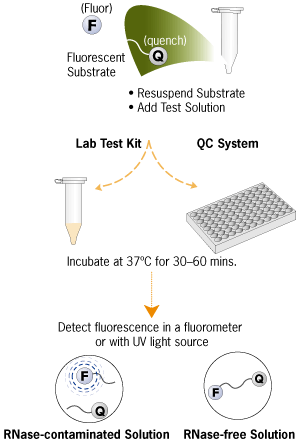
Figure 1. Schematic of RNaseAlert® Procedure.
RNaseAlert uses a novel RNA substrate tagged with a fluorescent reporter molecule (fluor) and a quencher to detect RNase activity. In the absence of RNases, the physical proximity of the quencher dampens fluorescence from the fluor. RNases cleave the RNA substrates, and the fluor becomes spatially separated in solution from the quencher, emitting a bright green signal when excited by light of the appropriate wavelength. Fluorescence can be readily detected by eye upon illumination on a UV box, or with a filter-based or monochromator-based fluorometer. Since the fluorescence of the RNaseAlert Substrate increases over time when RNase activity is present, results monitored with a fluorometer can be evaluated kinetically.
The RNaseAlert Lab Test Kit is formatted for small sample numbers, while the RNaseAlert QC System is designed for high throughput assays using 96 well plates.
- Sambashivan S, Liu Y, Sawaya MR, Gingery M, Eisenberg D (2005) Amyloid-like fibrils of ribonuclease A with three-dimensional domain-swapped and native-like structure. Nature 437(7056):266–9.
- Crestfield AM, Stein WH, Moore S (1963) Properties and conformation of the histidine residues at the active site of ribonuclease. J Biol Chem 228:2421–8.