Search
The RNase Protection Assay (RPA) is a widely used technique for the detection and quantitation of mRNA. The method involves:
- synthesis of a labeled antisense RNA probe complementary to a several-hundred-base region of the target mRNA,
- isolation of RNA sample(s) to be examined for target expression,
- hybridization of the labeled probe to a total RNA sample,
- treatment of the sample with single-strand-specific RNase to degrade unhybridized probe and target,
- separation of the remaining protected probe:target hybrids on a denaturing polyacrylamide gel
- detection/quantitation of the RNase-resistant "protected" probe.
The amount of protected probe is directly proportional to the amount of target mRNA in the sample. If an internal control probe or synthetic sense strand is included in the experiment, the assay can be used for relative and absolute quantitation, respectively.
The RPA is a more sensitive technique than Northern blotting. RPAs use solution hybridization, which is more efficient than traditional membrane-based hybridization, and RPAs can accommodate up to 100 µg of total or poly(A) RNA. Another advantage of RPAs over Northern analysis is that in RPAs, several RNA targets and internal controls can be assayed simultaneously.
Although mRNA analysis using reverse transcription followed by PCR (RT-PCR) is probably the most sensitive method for detecting low abundance targets, accurate quantitation can be difficult. Relative RT-PCR strategies require assessment in the exponential range of amplification, which limits sensitivity and can be difficult to optimize. Competitive RT-PCR requires synthesis of a carefully quantitated competitor and also requires significant amounts of optimization. Finally, real-time RT-PCR requires the use of special instrumentation that is not available in every lab.
The experiment described below demonstrates the use of Ambion's RPA III™ Kit (Cat #1414) to quantitate a target mRNA with a high degree of precision. It shows that this technique is capable of accurately detecting differences in target mRNA abundance that vary by as little as 20%.
Materials and Methods
The samples consisted of varying amounts of total RNA isolated from mouse liver using Ambion's RNAqueous™ Kit (Cat #1912). The amount of input mouse liver RNA in each hybridization reaction ranged from 0.5 µg to 20 µg. The appropriate amount of sheared yeast RNA (Cat #7118) was added so that each sample contained a total of 20 µg RNA. Samples were run in duplicate or triplicate to assess reproducibility.
Probe Synthesis
An antisense RNA probe to ß-actin was synthesized by in vitro transcription from a plasmid template containing a 245 bp fragment of the mouse cytoplasmic ß-actin coding region. The transcript was labeled by incorporation of 32P-UTP during synthesis.
The 20 µl transcription reaction contained 5 µl of 32P-UTP (10 mCi/ml, 800 Ci/mmol). No unlabeled UTP was added. The reaction was incubated for one hour at 37‘C and then treated with DNase I to degrade the plasmid template. Full-length transcript was purified on a denaturing polyacrylamide gel. The full-length probe was recovered from the gel slice by passive elution using the Elution Solution provided in the RPA III Kit.
The ß-actin probe (300 nt) contains the 245 nt coding region of ß-actin, as well as 55 additional bases from the vector polylinker region. The additional 55 bases are important because they allow the protected fragment to be distinguished from residual undigested full-length probe in the RPA.
Hybridization and Digestion Hybridization
reactions containing ~ 80,000 cpm of antisense ß-actin probe and target RNA as described above were ethanol precipitated and resuspended in a total volume of 10 µl RPA III Hybridization Solution. Samples were denatured for ~4 minutes at 95‘C and then hybridized overnight at 42‘C in a cabinet incubator. Following hybridization, excess probe was degraded by addition of 150 µl RPA III RNase Digestion Buffer (containing both RNase A and RNase T1) and incubation at 37‘C for 30 min. The RNase was then inactivated and the protected probe fragment recovered by addition of the RPA III RNase Inactivation/Precipitation Solution followed by centrifugation. Samples were resuspended in 8 µl of Gel Loading Buffer (supplied with the kit).
PAGE and Data Analysis
RPA samples were analyzed on a 5% polyacrylamide/8 M urea gel. The 12 x 14 cm gel was 0.5 mM thick and contained 20 wells. The gel was run at ~ 180 volts for 1.5 hours, then transferred to chromatography paper and dried under vacuum.
The amount of protected probe in each lane of the gel was quantified with a BioRad GS-250 phosphorimager. Data were plotted as pixel density (using the average pixel density value of replicate samples) vs input total mouse liver RNA.
Results
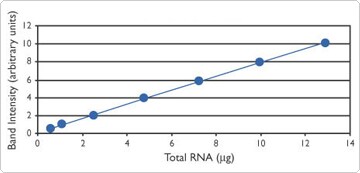
Using an RPA for mRNA Quantitation
The precision of an RPA is improved by assessing replicate samples (in duplicate or triplicate). When absolute quantitation of an mRNA target is needed, a standard curve can be constructed by performing the RPA on serial dilutions of a known amount of synthetic target (sense strand) side by side with the RPA of the unknown sample.
RPA III — A Simplified, Single-tube RPA
Ambion's RPA III Kit contains all the reagents and controls necessary for performing RPAs (except the antisense probe), and is designed to provide maximum sensitivity and convenience. New users will appreciate the detailed Instruction Manual and the inclusion of a positive control (probe template and target mouse liver total RNA) to validate performance of the kit. Perhaps the best advantage, however, is the kit's novel Inactivation/Precipitation Solution, which eliminates the need for proteinase K digestion and phenol extraction and makes the assay a single-tube procedure.