Search

Nuclease protection assays (NPAs), including both ribonuclease protection assays (RPAs) and S1 nuclease assays, are an extremely sensitive method for the detection, quantitation and mapping of specific RNAs in a complex mixture of total cellular RNA. The basis of NPAs is a solution hybridization of a single-stranded, discrete sized antisense probe(s) to an RNA sample (see Figure 1). The small volume solution hybridization is far more efficient than more common membrane-based hybridization, and can accommodate up to 100 µg of total or poly(A) RNA. After hybridization, any remaining unhybridized probe and sample RNA are removed by digestion with a mixture of nucleases. Then, in a single step reaction, the nucleases are inactivated and the remaining probe:target hybrids are precipitated. These products are separated on a denaturing polyacrylamide gel and are visualized by autoradiography. If nonisotopic probes are used, samples are visualized by transferring the gel to a membrane and performing secondary detection.
On this page
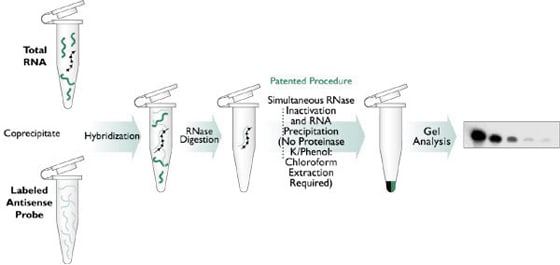
Figure 1. Detection of Specific mRNA Species Using a Nuclease Protection Assay.
NPAs are the method of choice for the simultaneous detection of several RNA species. During solution hybridization and subsequent analysis, individual probe/target interactions are completely independent of one another. Thus, several RNA targets and internal controls can be assayed simultaneously (up to twelve have been used in the same reaction), provided that the protected fragment of individual probes are of different lengths. NPAs are also commonly used to precisely map mRNA termini and intron/exon junctions.
RNA Quantitation
To quantitate mRNA levels using NPAs, the intensities of probe fragments protected by the sample RNA are compared to the intensities generated from either an endogenous internal control (relative quantitation) or known amounts of sense strand RNA (absolute quantitation). For more information on using NPAs for quantitation, see Technical Bulletin 151, "Use of Internal and External Standards or Reference RNAs for Accurate Quantitation of RNA Levels".
Advantages of NPAs over Northerns
- NPAs are more sensitive than traditional Northerns. They can be used to detect as little as 5 femtograms of target RNA or 4,000 to 50,000 copies/sample.
- NPAs are more tolerant of partially degraded RNA than Northerns. If samples are even slightly degraded, the quality of data from a Northern blot is severely compromised.
- NPAs are able to distinguish between transcripts of multi-gene families that may comigrate on Northerns.
- NPAs can be used to map mRNA termini and intron/exon junctions.
- Multi-probe assays are easy to perform with NPAs (see Figure 2). Up to 10 probes plus one or two internal controls can routinely be analyzed from a single RNA sample using Ambion's RPA III™ Kit. While multi-probe analysis is possible with Northern analysis it is very time consuming and requires multiple stripping and reprobing of a single blot.
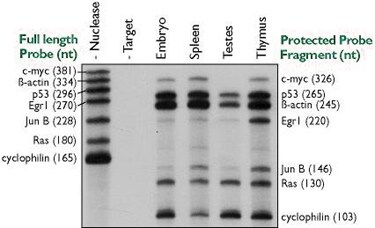
Figure 2. Simultaneous Quantitation of Multiple mRNAs Using the RPA IIIª Kit. Ten micrograms of various mouse tissue total RNAs were hybridized overnight with approximately 50,000 cpm each of seven distinct probe transcripts. Nuclease digestion, product separation on a denaturing 6% acrylamide gel, and a four-hour exposure to film at -80°C were used to assess hybridization levels. The radiolabeled probes were synthesized in 5 µl transcription reactions using Ambion's MAXIscript™ Kit with [-32P]UTP (800 Ci/mmol, 10 Ci/ml) and gel purified prior to hybridization. The specific activities of the cyclophilin and ß-actin probes were reduced twenty and 200-fold, respectively, by adding appropriate amounts of non-radioactive UTP to the transcription reaction.
Advantages of NPAs over RT-PCR Reactions
- RPAs are easy to use and don't require extensive optimization. Optimization reactions must be preformed prior to each RT-PCR reaction and the data obtained by RT-PCR analysis can be confusing and difficult to interpret.
- RPAs don't require expensive equipment purchases.
- Multi-probe assays are easy to perform with RPAs (see Figure 2). Up to 10 probes plus one or two internal controls can routinely be analyzed from a single RNA sample using Ambion's RPA III™ Kit. Due to interactions between primers, it is difficult to multiplex with greater than two primer sets using RT-PCR analysis.
- RPAs are able to distinguish between transcripts of multi-gene families that are not currently characterized.
Limitations of Nuclease Protection Assays
The primary limitation of NPAs is the lack of information on transcript size. The portion of probe homologous to target RNA determines the size of the protected fragment. Another drawback to NPAs is the lack of probe flexibility. The most common type of NPA, the ribonuclease protection assay, requires the use of RNA probes. Oligonucleotides and other single-stranded DNA probes can only be used in assays containing S1 nuclease. A region of the single-stranded, antisense probe must typically be completely homologous to target RNA to prevent cleavage of the probe:target hybrid by nuclease. This means that partially related sequences (e.g., probe and target RNA from different species) usually cannot be used.
Steps Involved in Nuclease Protection Assays
RNA Isolation
There are a number of protocols, techniques and commercially available kits that can be used to isolate RNA for NPAs, i.e., most if not all RNA isolation methods are compatible with NPAs. RNA isolation techniques all share these common attributes:
- Cellular lysis and membrane disruption
- Inhibition of ribonuclease activity
- Deproteinization
- Recovery of intact RNA
Ambion provides several options for isolation of total RNA and mRNA that are compatible with a variety of cells and tissues, including bacteria, yeast, plant and animal. For a further discussion of RNA isolation options, see RNA Isolation: The Basics.
Probe Generation and Purification
The type of probe used (RNA vs. DNA) is dependent upon which nuclease is used in the digestion step. Ribonuclease protection assays all require the use of RNA probes.
It is essential that probes used in NPA analysis are all of a discrete length. High specific activity single-stranded RNA probes can be produced by in vitro transcription reactions using Ambion's MAXIscript™ Kit. To assure that the probe is full length we recommend gel purification. Ambion's Technical Bulletin 171 discusses gel purification of probes in detail.
Either radiolabeled or nonisotopically labeled probes can be used for NPA analysis. We recommend that nonisotopic nucleotides be incorporated enzymatically (versus post synthesis chemical labeling of probes). Technical Bulletin 173, "Methods for Nonisotopic Labeling" describes how the MAXIscript Kit can be used to incorporate nonisotopic nucleotides into RNA probes.
For accurate quantitation of a specific message, probe concentration must be in molar excess over the target mRNA. This necessitates the use of low specific activity probes for abundant targets. For moderately abundant messages (e.g., ß-actin, GAPDH, or cyclophilin), a 1:50 dilution of labeled NTP with "cold" NTP should be used to increase the molar amount of probe made while simultaneously reducing the specific activity of the probe. For very abundant messages (e.g., 18S rRNA or 28S rRNA), a 1:10,000 dilution with "cold" NTP should be used.
The number and size of probes will dictate the gel size and acrylamide concentration. Typically a 5% denaturing acrylamide gel is used. This will effectively resolve fragments of about 500–1000 nucleotides. It is useful to have size markers on the gel. Single-stranded RNA markers are the most accurate (e.g., Ambion™ RNA Century-Plus Markers or Ambion™ Century Marker Templates), but double-stranded DNA markers can be used if it is not critical to know the exact size of the products.