Search
Isolating DNA and RNA Binding Proteins
Sequence-specific DNA and RNA binding proteins (e.g. promoters, gene regulatory proteins and transcription factors) are vital in normal cellular control and in disease states such as cancer and specific genetic diseases. These regulatory proteins are potential drug targets, but as these proteins are often short-lived and scarce you need a quick and sensitive method to purify them.
Choose Streptavidin-coupled Dynabeads for a simple, quick yet reliable way to purify sequence-specific DNA or RNA binding proteins.
Why choose streptavidin-coupled Dynabeads to isolate DNA/RNA binding proteins?
- Sensitive, rapid and convenient magnetic solid-phase handling
- Isolate pure and homogeneous transcription factors in 30
- Obtain high yields of proteins with only one adsorption step
- Enrich DNA and RNA binding proteins up to 20,000 fold
- DNA coupled to Dynabeads via the streptavidin-biotin association can be re-used at least ten times
The high stability and binding capacity of the DNA on Dynabeads allows binding of the target proteins with kinetics similar to that of DNA in free solution
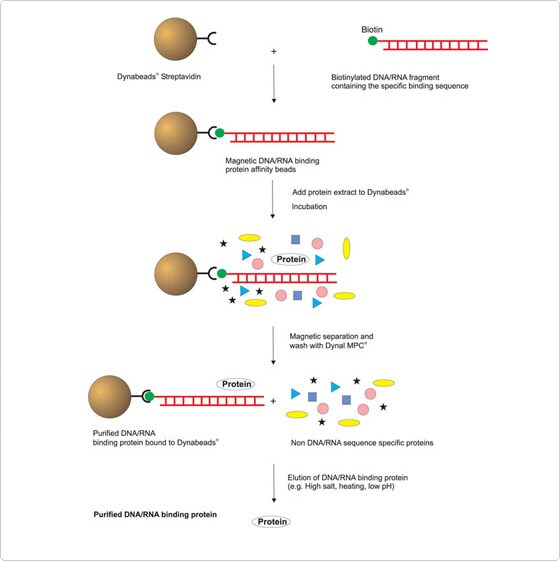
How does it work?
Immobilize the biotin-labelled DNA/RNA target sequence onto the beads and incubate with the cell extract. You can then isolate the bound proteins using magnetic separation. You can elute off the protein for characterization or apply the whole solid-phase complex directly e.g. to DNase footprinting studies.
- Dynabeads M-280 Streptavidin are ideal for isolating DNA and RNA binding proteins, due to the low non-specific binding of proteins to the beads.
- To immobilize larger DNA fragments, choose the Dynabeads kilobaseBINDER kit.
- Learn more about genome or gene expression analysis.
Selected References
- Mehta A. et.al. (1998) A sequence-specific RNA binding protein complements Apobec-1 to edit apolipoprotein B mRNA. Mol. Cel. Biol. 18(8):4426-4432.
- Nordhoff E. et al. (1999). Rapid identification of DNA-binding proteins by mass spectrometry. Nat. Biotechnol. 17: 884-888.
- Brodsky AS. and Silver A. (2002). A microbead-based system for identifying and characterizing RNA-protein interactions by flow cytometry. Mol. Cel. Proteomics 1(12):922-929.
- Gabrielsen OS et.al. (1989) Magnetic DNA affinity purification of yeast transcription factor t - a new purification principle for the ultrarapid isolation of near homogeneous factor. Nucl. Acids Res. 17:6253-6267
- Gabrielsen OS and Huet J. (1993) Magnetic DNA affinity purification of yeast transcription factor. Meth. in Enzym. 218:508-525.
- Ozyhar A. et.al. (1992) Magnetic DNA affinity purification of ecdysteroid receptor. J. Steroid Biochem. and Mol. Biol. 43(7):629-634 (read abstract)
- Ashley CT. et.al. (1993) FMR1 protein: conserved RNP family domains and selective RNA binding. Science 262:563-566 (read abstract)
- Mangiapan G. et al. (1996). Sequence capture-PCR improves detection of mycobacterial DNA in clinical specimens. J. Clin. Microbiol. 34(5):1209-1215
- Dong SM. et al. (2001). Detection of colorectal cancer in stool with the use of multiple genetic targets. J Natl Cancer Inst. 93(11): 858-865.
- Refseth UH. et al. (1997). Hybridization capture of microsatellites directly from genomic DNA. Electrophoresis. 18(9):1519-1523.
- Shuber AP. et al. (2002). Accurate, non-invasive detection of Helicobacter pylori DNA from stool samples: Potential usefulness for monitoring treatment. J. Clin. Microbiol. 40(1):262-264