Search
Combined isolation of cells and mRNA
This customer written article "Use of Immunomagnetic Separation, mRNA Isolation and RT-PCR for Detection of Activated Human Monocytes in Whole Blood" was published in DYNALogue 2/99 and is a good example of the combined use of cell-specific IMS followed by mRNA isolation.
Introduction

Blood monocytes are “blood stream passengers” en route to various tissue compartments where they are involved in numerous functions, including defence mechanisms, producing coagulation factors and cytokines, and phagocytosing various bacterial constituents and cellular debris (apoptotic bodies). These cells may be activated by a wide range of stimuli, such as immunological, inflammatory or bacterial components, to express a diversity of “early immediate genes”, including Tissue Factor (TF) and Tumour Necrosis Factor a (TNFa) (1-3). Combination of IMS (immunomagnetic separation) of monocytes from whole blood, mRNA isolation using Dynabeads Oligo (dT)25, reverse transcription (RT) and polymerase chain reaction (PCR) provides a powerful method for detecting such changes at the mRNA-level in activated monocytes.
The use of appropriate anticoagulants during blood collection is essential for obtaining reliable results. Heparin binds to, and activates, the naturally occurring clotting inhibitor, anti-thrombin. It will not bind calcium, which is required for initiation of the clotting cascade through the extrinsic and Tissue Factor dependent pathway. We have isolated monocytes from heparinised whole blood, stimulated in vitro with a component from gram negative bacteria, lipopolysaccharide (LPS), and investigated the recovery of monocytes after IMS and the LPS-induced gene-expression of TF- and TNFa-mRNA.
Materials and Methods
2 ml heparinised (15 U/ml) whole blood from healthy donors (n=5) were incubated for 3 hours at 37°C, rotated (50 rpm in a GFL 3032 Incubator) in the presence or absence of 2 ng/ml LPS (E. coli Endotoxin, BioWhittaker). Following incubation, the blood was chilled on ice-water for 10 minutes to reduce the monocytes ability to phagocytose the beads. The ice cold blood (250 µl) was then subjected to IMS using 25 µl (4 x 108 beads/ml) Dynabeads CD14 to isolate monocytes.
The blood and beads were mixed and incubated at 4°C for 10 minutes in a Dynal Sample Mixer. The Dynabeads-monocytes complexes were isolated with a magnet (Dynal MPC). The number of monocytes in the whole blood was counted before and after IMS in a Technicon H.2TM System. The separated monocytes were washed four times in ice cold PBS before being lysed in 250 µl lysis buffer. The beads were then separated from the lysate using the Dynal MPC magnet, and the lysate either frozen at -70°C or immediately subjected to mRNA isolation.
200 µl lysate was incubated with 25 µl (3.3 x 108 beads/ml) Dynabeads Oligo (dT)25 (prewashed and dispersed in 200 µl lysis buffer) in a sample mixer for 5 minutes at room temperature. After four wash steps, using the Dynal MPC between each step, the beads were resuspended in 50 µl 10mM TrisHCl and 2 µl of the Dynabeads-mRNA complexes were subjected to a two-step RT-PCR in a GeneAmp PCR System 2400 (Perkin Elmer).
A RT-mix (18 µl) containing 5 mM MgCl2, 1 x PCR buffer II, 1 mM each of dATP, dCTP, dGTP and dTTP, 1 U RNase Inhibitor (all reagents purchased from Perkin Elmer), 16 U M-MuLV Reverse Transcriptase (FPLC pure, Amersham/Pharmacia Biotech) and DEPC treated water was added to the Dynabeads-mRNA complexes. The RT-step was performed at 37°C for 60 minutes, at 99°C for 5 minutes and then stored at 4°C. The PCR-step was performed by adding 80 µl PCR-mixture containing 1.5 mM (TF) or 2.0 mM (TNFa, b-actin) MgCl2, 1 x PCR buffer II, 2.5 U AmpliTaq Gold (Perkin Elmer) and 25 pmol of each of the primers. The TF primers annealed at 55°C and the TNFa and b-actin primers at 60°C.
To normalise the measurements in different samples due to variations in the isolation steps (IMS and mRNA), the expression of the housekeeping-gene, b-actin, was quantified (4).
The b-actin-mRNA expression in the LPS treated samples was set to 100% and the other samples adjusted accordingly.
To ensure that the PCR-products were generated from cDNA, we used primers that either generated larger amplification products from genomic DNA (TF and b-actin) or did not amplify genomic DNA at all (TNFa). TF-mRNA, TNFa-mRNA and b-actin-mRNA were analysed after pilot studies to determine the optimal number of PCR-cycles to be within the linear range of amplification for both the gene of interest and the housekeeping gene. Primers used are listed in the margin.
The PCR products were examined by agarose 3 w/v % gel (stained with ethidium bromide) electrophoresis (100 V, 30 minutes) and the fluorescence detected by Kodak Image Station 440 CF (Kodak Digital Science TM 1D Image Analysis Software). Amplisize TM DNA Size Standard 50-2000 bp ladder (Bio-Rad Laboratories) was used as a molecular weight standard.
Results
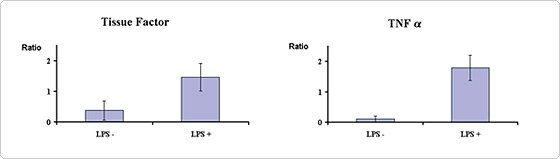
IMS of monocytes in whole blood resulted in 92 % depletion (n=3). LPS incubation of whole blood prior to IMS resulted in increased amounts of TF- and TNFa-mRNA in isolated monocytes.
Expressed as ratios, the non LPS incubated blood gave: TF-mRNA/b-actin-mRNA = 0.38 (Standard Error of the Mean, SEM:0.3) and TNFa-mRNA/ b-actin-mRNA = 0.115 (SEM: 0.099) (Figure 2). LPS incubation resulted in TF-mRNA/b-actin-mRNA = 1.47 (SEM:0.44) and TNFa-mRNA/b-actin-mRNA = 1.795 (SEM:0.424).
Concluding Remarks
It is well documented that LPS induces expression of TF-mRNA and TNFa-mRNA in cultured human monocytes (5). Here we have shown that human monocytes stimulated with LPS in whole blood, conditions more similar to the in vivo situation, also demonstrate increased expression of TF-mRNA and TNFa-mRNA.
IMS combined with mRNA-isolation and RT-PCR is a powerful method to detect changes at the mRNA-level in human blood cells. This approach lends itself to the rapid examination of gene expression in these cells both under normal circumstances and during disease.
By using this approach for sequential monitoring of mRNA expression of various cytokines (eg. TNFa, IL 6, IL 8) and coagulation factors (eg TF) in monocytes obtained from patients with infections (6) or unstable angina (7), our understanding of important cell functions may be increased.
The following primers were used:
- TF 5’primer: 5’CTC GGA CAG CCA ACA ATT CAG AGT 3'.
- TF 3’primer: 3' TGT TCG GGA GGG AAT CAC TGC TTG AAC ACT 5'. From Medprobe, expected product sizes are 270 bp (cDNA) and 883 bp (genomic).
- TNFa 5’primer: 5’CAG AGG GAA GAG TTC CCC AG 3'.
- TNFa 3’primer: 3’CCT TGG TCT GGT AGG AGA CG 5'. From PE-Applied Biosystems UK, expected product size is 325 bp (cDNA).
- b-actin 5’primer: 5’TGA CGG GGT CAC CCA CAC TGT GCC CAT CTA 3'.
- b-actin 3’primer: 3’CTA GAA GCA TTT GCG GTG GAC GAT GGA GGG. 5'. From Stratagene (cat 030210), expected product sizes are 661 bp (cDNA), 876 bp (genomic).
References
- Kirchhofer D, Nemerson Y. Initiation of bloodcoagulation: the tissue factor/factor VIIa complex. Curr.Opin.Biotechnol. 1996;7(4):386-91.
- Mackman N. Regulation of tissue factor gene expression in human monocytic and endothelial cells. Haemostasis 1996;26 Suppl 1:17-19.
- Osnes LT et al. Lipopolysaccharide activation of human monocytes mediated by CD14, results in a coordinated synthesis of tissue factor, TNFa and IL-6. J.Endotox Res. 1995;2:27-35.
- Soutar RL, Dillon J, Ralston SH. Control genes for reverse-transcription-polymerase chain reaction: Br.J.Haematol. 1997;97(1):247-8.
- Osnes LT et al. Acid and sodium salicylate inhibit LPS-induced NF-kappa B/c-Rel nuclear translocation, and synthesis of tissue factor (TF) and tumor necrosis factor alfa (TNF-alpha) in human monocytes. Thromb.Haemost. 1996;76(6):970-6.
- Rivers RP, Cattermole HE, Wright I. The expression of surface tissue factor apoprotein by blood monocytes in the course of infections in early infancy. Pediatr.Res. 1992;31(6):567-73.
- Leatham EW, Bath PM, Tooze JA, Camm AJ. Increased monocyte tissue factor expression in coronary disease. Br.Heart J. 1995;73(1):10-13.
Article written by Reidun Øvstebø, Kari Bente Foss Haug, Liv Osnes, Gun Britt Joø, Peter Kierulf. The Research and Development Group, Clinical Chemistry Department, Ullevål University Hospital, Oslo, Norway.