Search
High Throughput RNA Isolation: Methods Comparison
Introduction
The demand for high throughput RNA isolation has been dramatically increasing with wide application of RNAi, expression profiling and molecular diagnosis. We will present a comparison of various RNA isolation methods that have been adapted to high throughput platforms, focusing on consistently high yield and quality of isolated RNA, reduction of cross-contamination, and simplicity and robustness of the protocol. 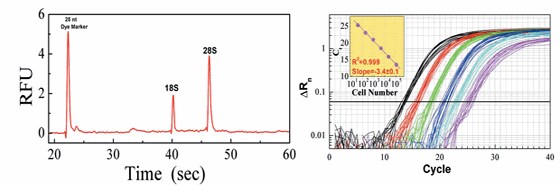
Figure 1. RNAqueous®-96 Filter Plate-Based RNA Isolation. K562 cells were used. Panel A shows the high quality of isolated RNA. Panel B shows the RNA linear recovery exemplified by GAPDH mRNA quantified by qRT-PCR.
RNA Isolation with Filter Plates
RNA Isolation with Magnetic Beads
The glass fiber filter based RNA isolation method is simple and easy to use. However, there are some potential problems: 1) filters may clog with tissue samples or high cell input; 2) the plate may fail to seal properly with the vacuum manifold, thus a robust protocol often requires a centrifuge; 3) RNA may not be efficiently recovered when sample input is very low. To address all these problems, we have developed a new method that uses magnetic microspheric beads to selectively bind nucleic acids. In general, a microspheric bead-based approach results in more consistent RNA recovery and more thorough DNA treatment than glass fiber filter based RNA methods. This is because beads can be fully resuspended in solution to enable more thorough mixing, washing and elution. In addition, bead-based methods are easier to automate than filter plate-based methods where vacuum may cause cross-contamination and the occasional filter clog may cause isolation failure of all samples in the whole plate. We will show comparative results on RNA isolated from cultured cells, as well as for viral RNA isolated from liquid biological samples.
Solid phase RNA isolation with magnetic beads is very suitable for high throughput applications, and the process is easy to automate. Advantages of Ambion’s new RNAqueous-MAG technology include:
- high purity, integrity and yield with high consistency
- 1-1.5 hours to process up to 4 plates in parallel
- RNA eluted in 20-50 µl water for easy integration with qRT-PCR or RNA amplification
This protocol is routinely used at Ambion for siRNA evaluation. It has also been successfully adapted for high throughput viral RNA isolation for molecular diagnosis. 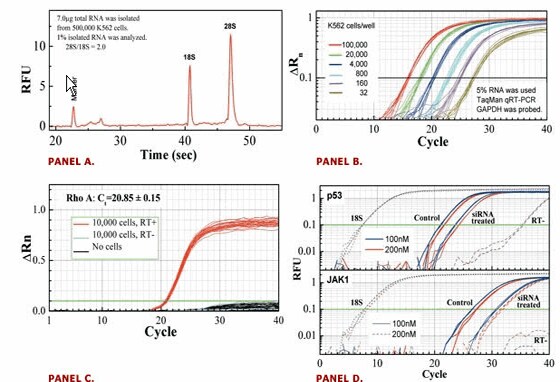
![]()
Figure 2. RNAqueous-MAG Performance and Application. Panel A shows high integrity and quality of isolated RNA; Panel B shows the RNA linear recovery; Panel C shows no cross contamination and efficiency in DNA removal. Panel D shows its application in siRNA evaluation (Human umbilical vein endothelial cells were transfected and mRNA was isolated 24 hr after transfection and quantified by qRT-PCR with SYBG.)
| Cell Type | RNA yield* (pg/cell) |
| K562 | 25 |
| HeLa S3 | 12 |
| BJ (human primary) | 10 |
| CHO (hamster) | 9.2 |
| COS-7 (monkey) | 10 |
| NIH/3T3 (mouse) | 10 |
| MCF-7 | 16 |
Skip RNA Isolation for qRT-PCR
High quality RNA is critical for global gene expression profiling with microarrays. On the other hand, for expression profiling of a handful genes using TaqMan® real-time RT-PCR, purified RNA may not be necessary. The real-time procedure produces excellent results due to the high specificity derived from gene specific primers and probes, as long as nucleases and potential polymerase inhibitors are deactivated. Ambion’s Cells-to-cDNA™ technology (US patent pending), by taking full advantage of the high specificity of TaqMan qRT-PCR, completely bypasses RNA isolation thereby increasing the throughput potential, improving the precision of the final results and decreasing overall costs. It is suitable for quantifying abundant and moderately expressed mRNAs. Simply add lysis solution and heat for 15 min. The lysate is ready for a two-step RT-PCR or coupled one-step RT-PCR. 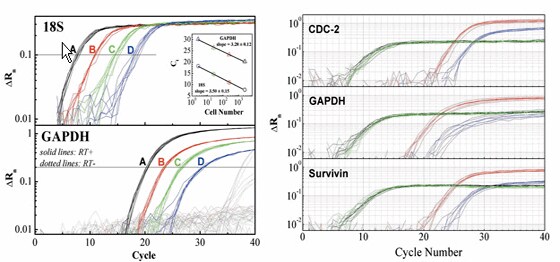
Figure 3. mRNA Quantification Without RNA Isolation by Ambion’s Cells-to-cDNA Technology. Left Panel: the linearity for 100 to 100,000 HeLa cells input by qRT-PCR. Right Panel: siRNA validation by qRT-PCR using Cells-to-cDNA technology. After siRNA transfection, target gene expression was down by ~ 10 fold. 18S was chosen for normalization of cell input.