Search
Cells-to-cDNA II Using FACS Sorted Cells
Introduction
Fluorescence activated cell sorting (FACS) is a tool for measuring phenotypic, biochemical and molecular properties of individual cells using fluorescent probes that bind specifically to molecules associated with the cell. This technology is commonly used to collect cells of a given population down to the level of a single cell.
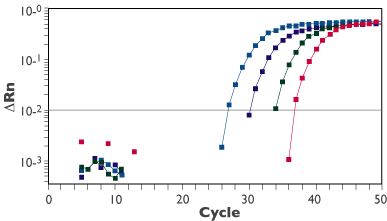
Figure 1. Linear Amplification from 1-1000 Cells.
Dr. Daniel Douek of the NIH Vaccine Research Center recently used Ambion's Cells-to-cDNA™ II Lysis Buffer to analyze gene expression of human ß-glucuronidase (hGUS) in FACS sorted Jurkat cells using real-time RT-PCR. GUS is an enzyme constitutively expressed at moderate levels in all cells.
Jurkat cells were sorted by FACS and collected at 1, 10, 100 and 1000 cells per well in a 96-well PCR plate. Each well was preloaded with 4 µl of Cells-to-cDNA II Lysis Buffer. After the cells were sorted and lysed, the plate was placed at -80°C for 1 week. The lysates were then DNase treated according to the Cells-to-cDNA II protocol. One-step RT-PCR was performed in 20 µl using a TaqMan® probe for hGUS. Six replicates were run for each cell concentration. A representative amplification plot is shown in Figure 1. Results show quantitative amplification from 1-1000 cells, demonstrating a 3-log dynamic range and reproducibility of the assay.