Search
How to Conduct Fragment Analysis
Fragment analysis is a technique for separating DNA molecules by size that can be performed using capillary electrophoresis (CE). In contrast to Sanger sequencing, which yields information about the identity and order of the four nucleotide bases in a segment of DNA, fragment analysis assigns a length in bases to individual DNA fragments. Fragment analysis is a lesser-known CE method, but it has significant advantages, including multiplexing, for applications requiring size, relative quantitation, and genotype information.
The DNA fragment analysis workflow consists of four general steps: DNA extraction, PCR amplification, capillary electrophoresis, and data analysis (Figure 1). The workflow begins with the isolation, purification, and quantitation of the DNA that will be used to generate labeled PCR amplicons. The amplicons are then separated by size during capillary electrophoresis, and the fragments are visualized in data analysis as peaks corresponding to alleles.

Primer design
Two primers are used for fragment analysis: one to bind to the 5' end of the template gene and a second to bind to the 3' end. One of the primers is labeled with a fluorescent molecule (i.e., a fluorophore) at the 5' end. If multiplexing, primers with the same fluorophore should not be used that produce fragments of a similar length. The resolution and number of targets can be increased by spreading different colored fluorophores across the available size range (usually 50–1,200 bp).
It is important to design primers that will bind to their target sequences during thermal cycling (for general guidelines on primer design, see [1]). There are several free online tools that can assist with primer design.
DNA extraction
The quality of the DNA fragments to be analyzed can be significantly affected by the characteristics of the sample itself and the method chosen for nucleic acid extraction and purification. Ideal methods will vary depending on the source or tissue type, how the sample was obtained from its source, and how it was handled or stored prior to extraction. Common sources include tissue, blood or plasma, cells, and plant material. DNA can be extracted using a kit or following a lab-developed protocol.
PCR amplification
Next, primers are used to amplify the extracted DNA via PCR.
There are several PCR master mixes available that contain a high-performance DNA polymerase, salts such as magnesium chloride (MgCl2), buffer, and nucleotides, to simplify setting up PCR reactions. The other reaction components required are template DNA, primers, and nuclease-free water.
Once the reactions are set up, they are run on any thermal cycler. A typical PCR program includes initial denaturation followed by repeated cycles of denaturation, annealing, and extension steps, with a final hold at 4°C. The annealing temperature should be optimized empirically.
Capillary electrophoresis
Before the samples are run on a genetic analyzer, a plate is prepared containing formamide and a set of fluorescently labeled DNA fragments of known length to act as a size standard. The size standard should span the expected range of fragments in your experiment. Ideally, it should contain two fragments above and two fragments below the size of each target fragment. There are several commercially available size standards.
Each sample is run in a dedicated glass capillary as part of an array. The fragments enter the capillary as a result of electrokinetic injection. A high-voltage charge applied to the buffered sequencing reaction forces the negatively charged DNA fragments into the capillaries. The DNA fragments are separated by size due to the larger fragments migrating more slowly through the matrix. CE can separate the fragments with single-base resolution.
Data analysis
As the fragments migrate through the capillary, a laser excites the fluorescent dye labels, and the signals are displayed as peaks (Figure 2).
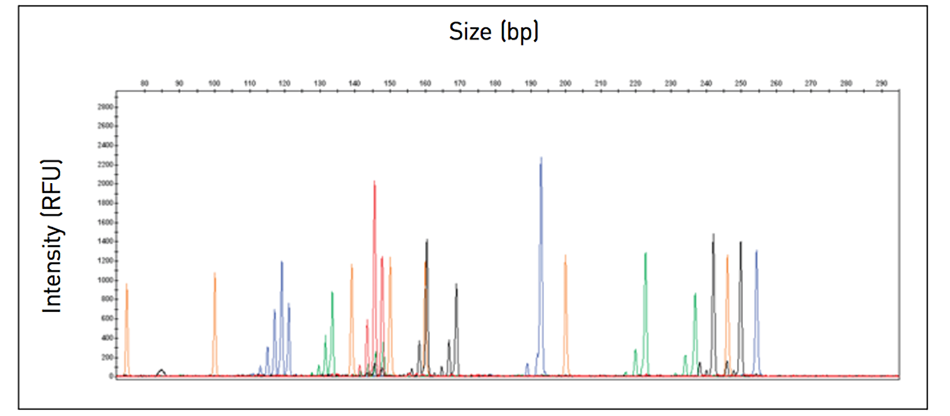
Figure 2. In fragment analysis, fluorescently labeled fragments are separated and sized according to an internal standard.
Data analysis software can provide a profile of the separation, precisely calculate the sizes of the fragments, and determine the alleles present in the sample.
Microsatellite analysis
Microsatellites are highly variable short tandem repeats (STRs) of genomic DNA that can be used to distinguish between individuals by fragment analysis. Their diversity makes them ideal for population genetics studies, use in forensic casework, or to establish paternity. The CLA Identifiler Direct PCR Amplification Kit amplifies DNA from 16 different human microsatellite loci, generating allelic data that can be used to uniquely identify a specimen. Applications include:
- Cell line authentication—There have been a large number of cases where the identity of cell lines given in publications are later shown to be incorrect. The Identifiler kit can be used to demonstrate that cell lines used are authentic before, during, and after experiments.
- Mixed sample analysis—Any experiment that involves tracking unique human cells being mixed with populations of other cells, human or otherwise, might benefit from establishing identity. One example of this is human tumors being implanted in mouse models for study; the xenografts can be tested to authenticate the work.
- Forensics— Microsatellite analysis is a primary tool used by the forensic community and has a well-established track record.
Reference
- Lorenz TC (2012) Polymerase Chain Reaction: Basic Protocol Plus Troubleshooting and Optimization Strategies. J Vis Exper 63:e3998.
For Research Use Only. Not for use in diagnostic procedures.