Search
From Exosome Isolation to Biomarker Discovery
Total Exosome Isolation Reagents for a Complete Exosome Workflow
(See a complete list of products discussed in this article.)
Exosomes are small (30–150 nm) vesicles secreted by mammalian cells that, along with their unique RNA and protein cargo, have become the focus of much research. Critical to furthering our understanding of the function of exosomes and their applications in diagnostics and therapeutics is the availability of reagents and protocols for their isolation and characterization, as well as for the analysis of their RNA and protein content. We have developed a new set of reagents that enables a complete exosome workflow—from the fast and efficient extraction of exosomes from cell culture medium or serum to the isolation and characterization of exosomal RNA content using next-generation sequencing.
Small Vesicles With Huge Potential
Exosomes are small vesicles thought to comprise RNA and protein molecules that are secreted by all mammalian cell types in culture, and also occur naturally in body fluids including blood, saliva, urine, and breast milk. The precise molecular mechanism of their secretion and uptake, as well as their composition, cargo, and functions, are only beginning to be unraveled [1]. Originally thought of as “garbage bags” used by cells to get rid of unnecessary macromolecules, exosomes are now viewed as specifically secreted vesicles that enable intercellular communication [2]. Studies have shown that exosomes can transport mRNA and miRNA between cells, and that the transferred RNA is functional in its new location [3].
Total Exosome Isolation Reagents
Isolation of exosomes from cell culture medium and body fluids is presently a tedious, difficult, and nonspecific process that requires ultracentrifugation [4]. To simplify and shorten the process of exosome isolation, we have developed two different Total Exosome Isolation Reagents that enable straightforward and reliable concentration of intact exosomes from cell culture medium or blood serum. By tying up water molecules, the reagents force less-soluble components (i.e., exosomes) out of solution, allowing them to be collected after a brief, low-speed centrifugation step. The procedure enables recovery of a significant number of microvesicles in the typical exosome size range of 30–150 nm, with size distribution comparable to that of exosomes isolated using ultracentrifugation (Figure 1).
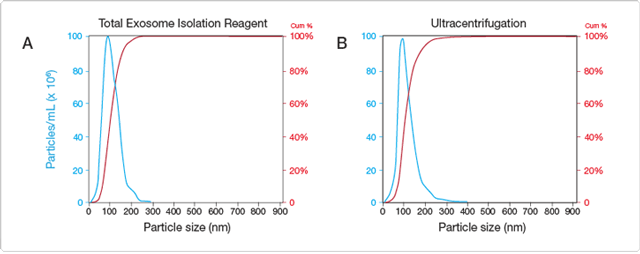
Figure 1. Analysis of exosomes recovered from human blood serum using the Total Exosome Isolation Reagent or an ultracentrifugation protocol. Exosomes were extracted from human blood serum using (A) Total Exosome Isolation Reagent or (B) a “homebrew” ultracentrifugation procedure [4], and analyzed on the NanoSight® LM10 instrument. Both protocols yield exosomes with comparable size distribution: all nanoparticles are smaller than 300 nm, with the majority ~50–150 nm in size.
The Total Exosome RNA and Protein Isolation Kit
The Total Exosome RNA and Protein Isolation Kit was designed for the isolation of RNA and protein from an enriched exosome preparation. A resuspended exosome preparation is divided into two samples; one is used for common protein analysis methods such as western blotting, and the other is used for organic extraction followed by glass-fiber filter purification of total RNA. Maximal yields of ultrapure RNA can be prepared in about 30 minutes and are suitable for studies of RNA expression (including miRNA), processing, or function. The RNA can also be used in a variety of downstream applications including RT-PCR, sequencing (e.g., using an Ion Personal Genome Machine® (PGM™), Ion Proton™, or SOLiD® System), RNA amplification, microarray analyses, and solution and blot hybridization.
Sequencing and Analysis of Exosomal RNA
Exosomal RNA extracted using the Total Exosome Isolation Reagent protocol can be analyzed by sequencing and then compared to RNA from the “parental” samples (cells or serum from which the exosomes originated). Small RNA libraries can be prepared using the Ion Total RNA-Seq Kit v2, with some protocol modifications to address the relatively low amounts and small sizes (<200 nt) of exosomal RNA. Once the libraries are constructed, they can be sequenced using the Ion PGM™ Sequencer and standard protocols.
After sequencing, we compared the RNA content of exosomes extracted from HeLa cell culture medium and from serum to that of their respective parental samples (HeLa cells and whole serum). The exosomes were found to contain a diverse population of RNA, including miRNA, mRNA, rRNA, and other small RNA (snoRNA and other ncRNA). Overall, exosomal cargo for both sample types reflects the RNA content of the parental cells—in terms of sequences and expression levels—for many targets (Figure 2), suggesting that exosomal RNA could serve as an important diagnostic tool. Nevertheless, certain miRNA, tRNA, ncRNA, and mRNA show significant differences in levels between the exosomal and parental samples (Figure 3), which may lead to the identification of exosome-specific markers.
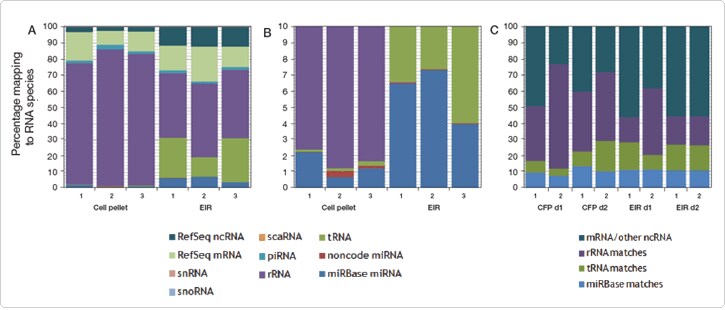 Figure 2. Distribution of RNA species from HeLa cell and human serum exosomes and parental samples.(A) Distribution of RNA species derived from HeLa exosome preparations obtained with the Total Exosome Isolation Reagent (EIR) and parental samples (cell pellet) at full scale. (B) The data in (A) zoomed in to the 0–10% distribution range to view less prominent RNA types and miRNA representation. (C) Distribution of RNA species derived from the serum exosome preparation (EIR) and parental cell-free plasma sample (CFP) using blood serum from two human donors, d1 and d2. RNA was sequenced using the Ion Personal Genome Machine® (PGM™) Sequencer. Reads were mapped to miRBase microRNA precursors (hairpins), and remaining unmapped reads were mapped to NCBI RefSeq transcripts and then to the Noncode.org noncoding RNA database.
Figure 2. Distribution of RNA species from HeLa cell and human serum exosomes and parental samples.(A) Distribution of RNA species derived from HeLa exosome preparations obtained with the Total Exosome Isolation Reagent (EIR) and parental samples (cell pellet) at full scale. (B) The data in (A) zoomed in to the 0–10% distribution range to view less prominent RNA types and miRNA representation. (C) Distribution of RNA species derived from the serum exosome preparation (EIR) and parental cell-free plasma sample (CFP) using blood serum from two human donors, d1 and d2. RNA was sequenced using the Ion Personal Genome Machine® (PGM™) Sequencer. Reads were mapped to miRBase microRNA precursors (hairpins), and remaining unmapped reads were mapped to NCBI RefSeq transcripts and then to the Noncode.org noncoding RNA database.
 | Figure 3. Differential RNA representation between exosome and parental samples. Specific serum-derived miRNAs from cell-free plasma (CFP) and Total Exosome Isolation Reagent (EIR) samples. Quantities were normalized by the percentage of reads aligned to that transcript out of the total mapped. |
References
- Vlassov AV, Magdaleno S, Setterquist R et al. (2012) Biochim Biophys Acta 1820:940–948.
- Mittelbrunn M, Sánchez-Madrid F (2012) Nat Rev Mol Cell Biol 13:328–335.
- Valadi H, Ekström K, Bossios A et al. (2007) Nat Cell Biol 9:654–659.
- Théry C, Amigorena S, Raposo G et al. (2006) Curr Protoc Cell Biol Chapter 3: Unit 3.22.
Resources | |
| Article Download Get a copy of this article as it appears in the print version of BioProbes 68. | Learn More About: |
FOR RESEARCH USE ONLY. NOT FOR USE IN DIAGNOSTIC PROCEDURES.