Search
Absolute Quantitation of mRNA Species using Northern Blotting, Nuclease Protection Assays and RT-PCR
Absolute quantitation measures the absolute amount (e.g. 5.3 x 105 copies) of a specific mRNA in an experimental sample. Dilutions of a synthetic sense strand RNA (identical or very similar to the target template) are co-amplified or detected along with the endogenous target. The sense RNA creates a concentration curve to which the endogenously expressed message is then compared in order to obtain an absolute measurement of the transcript under study. Here, we discuss the use of such synthetic sense-strand RNAs for absolute quantitation in Northern blotting, nuclease protection assays, and RT-PCR.
Absolute quantitation using Northern analysis
While Northern blotting is widely perceived as being a valuable qualitative assay to obtain information about mRNA expression, absolute quantitation by Northern analysis is both possible and straightforward. A sense strand RNA transcript complimentary to the probe is synthesized. This synthetic RNA is quantitated, and dilutions are spiked into a yeast RNA background and size fractionated in a denaturing agarose gel next to the sample RNAs. Alternatively, a longer or shorter sense RNA, sharing target sequence with the endogenous message, can be added to the experimental samples. After blotting, the membrane is probed with an antisense probe and the amount of endogenous target is compared with the concentration curve generated by the synthetic sense strand dilutions.
Figure 1 illustrates analysis of ß-actin expression in various mouse tissues. For example, by comparing the expression of ß-actin in mouse liver to the standard curve derived in this experiment, it demonstrates that there are approximately 30 pg of ß-actin RNA/µg of total mouse liver RNA.

Figure 1. Quantitation of variable ß-actin expression in mouse tissues.Mouse total RNA from several tissues was run on a Northern gel and transferred to membrane using the Ambion™ NorthernMax™ Kit. Dilutions of full-length ß-actin sense RNA was spiked into a yeast background and also run on the same gel. A 32P-labeled antisense ß-actin probe was used to detect ß-actin mRNA in the mouse tissues as well as the artificial sense strand. Absolute expression levels were determined by direct comparison of the signal from the mouse tissues with the concentration curve generated with artificial sense strand ß-actin.
Absolute quantitation of mRNA species using nuclease protection assays (NPA)
Different procedures exist for determining the abundance of a particular mRNA in a heterologous RNA sample using nuclease protection assays. Like in the Northern blotting example above, a standard curve is constructed using known amounts of in vitro synthesized "sense strand RNA" hybridized with an excess of labeled antisense probe (See Figure 2). Experimental samples are analyzed in conjunction with the control reactions containing a titration of the unlabeled (or labeled to low specific activity) sense strand RNA used to generate the standard curve. The intensity of the probe fragments protected by the experimental RNA samples can then be compared directly to the standard curve and used to define the absolute amount of the target RNA species present in the RNA samples.
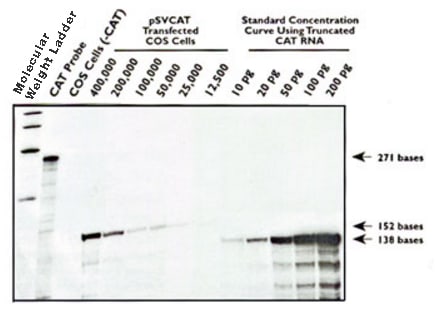
Figure 2. Detection of chloramphenicol acetyltransferase (CAT) expression in COS Cells transiently transfected with pSVCAT. Increasing dilutions of pSVCAT-transfected COS cells were assayed for CAT mRNA by ribonuclease protection. Reaction products were resolved on a denaturing 8% polyacrylamide gel, dried, and autoradiographed. Lane 1, Molecular weight ladder; lane 2 undigested CAT probe; lane 3, CAT probe digested after hybridization with total RNA from 400,000 COS cells; lanes 4-9, CAT probe digested after hybridization with total RNA from 400,000, 200,000, 100,000, 50,000, 25,000, and 12,500 transfected COS cells; lanes 10-14, standard concentration curve generated by digestion of the CAT probe after hybridization to 10, 20, 50, 100, and 200 pg of truncated sense transcript diluted in 5 µg of yeast RNA.
Synthesis of "sense strand RNA"
The synthetic sense strand transcripts used for absolute quantitation of Northern blots and nuclease protection assays can be made by linearizing the DNA template used for antisense probe synthesis on the other side of the probe insert, and synthesizing transcript from the opposite strand, provided that the insert is flanked by opposable promoters. Use of an internal restriction site for linearization will generate a "truncated" sense strand. This may be desirable when the number of experimental samples is large, since the concentration curve can then be added into the experimental samples; i.e.. both natural sense strand and artificial sense strand can be in the same tube.
Synthesis is usually performed in the presence of all four unlabeled rNTPs (for RNA probes synthesized in in vitro transcription reactions). Under these conditions, a large mass of sense strand is made. A tracer amount of labeled nucleotide is sometimes added to the four unlabeled ribonucleotides to aid in calibration (see below). Sense strand made in this way with or without label will be stable for at least several months. The Invtrogen™ Ambion™ MEGAscript™ and MAXIscript™ Kits are ideal for this purpose.
Gel Purification?
Sense strand molecules can be gel-purified. However, since the sense strand is typically being synthesized in the presence of high concentrations of all four rNTPs, transcripts should be mostly full-length, and gel purification may therefore not be necessary. DNase I can be used to degrade the DNA template. In any case, it is at least a good idea to check the integrity of the sense strand transcript on a gel before use. If gel purification is desirable, the position of unlabeled transcripts can be determined either by:
- Running a sample of the same-sized labeled antisense strand probe in a neighboring lane and using its position;
- UV shadowing of the bands; or
- Staining with EtBr or acridine orange.
Note that if dyes are used, they must be removed after gel elution, since they can compromise hybridization to the sense strand. UV shadowing involves placing the gel on a fluor-coated TLC plate (Cat. # 10110G) and shining short wavelength (254 nm) UV light onto the gel. Nucleic acids will show up as purple bands. While this method requires no staining, it does require a minimum of approximately 0.3-0.5 µg nucleic acid.
Calibration of the sense strand
Exact quantitation of the sense strand transcript is important to generate an accurate curve. The sense strand can be quantitated by reading absorbance at OD260 or by calculating yield based on the percent incorporation of a radiolabeled tracer. 3H-nucleotides can be used for this purpose although a small amount of 32P, 33P or 35S radiolabeled nucleotides (e.g. 1 µl of a 1:10 dilution of the stock labeled nucleotide, added to the synthesis reaction in addition to the unlabeled form of this nucleotide) will also work.
Absolute or competitive RT-PCR
Competitive RT-PCR precisely quantitates a message by comparing signal intensity of the RT-PCR product to a concentration curve created using a synthetic transcript or "competitor". Competitors are designed as follows:
- To be amplified using the same primers as the desired target sequence,
- To have the same amplification efficiency as the target sequence,
- To be of a significantly different size from the target to allow differentiation of the two products on an agarose gel, and
- to control for variations in the RT reaction.
Thermo Fisher Scientific has two products that help researchers meet all of these criteria: the RT-PCR Competitor Construction Kit and the Competitive Quantitative RT-PCR Kits. Competitor RNA transcript is synthesized, quantitated and diluted into the sample RNA. Pilot experiments are run to determine the range of competitor concentration where the experimental signal is similar in concentration to the endogenous target. A more precise concentration can then be determined by repeating the experiment with a smaller dilution range, generating data that approximates abundance of the endogenous target in the sample.
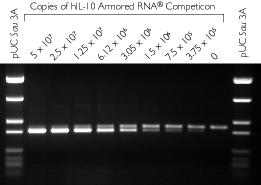
From the data illustrated in Figure 3 it can be determined that between 1.5 - 3.05 x 106 copies of hIL-10 RNA are present in the sample.
Products available
Thermo Fisher Scientific offers a variety of internal standards as linearized plasmid templates to generate antisense RNA or DNA probes by in vitro transcription or primer extension. These include human, rat and mouse templates for ß-actin, GAPDH and cyclophilin and regions of 18S and 28S rRNA that cross react with most vertebrates with few, if any, mismatches (the Ambion™ 18S rRNA template can be used with virtually any eukaryote). All transcription templates (except pT7-18S-rRNA) are inserted into our unique pTRIPLEscript™ vectors. pTRIPLEscript™ vectors feature three tandem phage promoters which offer the researcher the convenience of synthesizing an antisense RNA transcript using either SP6, T7 or T3 polymerase. They are ideal for the synthesis of antisense internal control standards and can be used in Northern blots or as a second probe in ribonuclease and S1 protection assays for simultaneous detection of both an internal reference RNA and the mRNA being studied. Thermo Fisher Scientific also provides DECAtemplates™ for making random-primed DNA probes for commonly used internal controls (18S rRNA, ß-actin, GAPDH, and cyclophilin).
Along with our line of internal control templates, we offer an extensive line of transcription templates for human and mouse oncogene studies as well as high quality mouse total RNA for RNA expression analysis. The Ambion™ MAXIscript™ Kits are ideal for the transcription of high specific activity antisense probes (the Ambion™ MEGAshortscript™ Kits are another option for transcribing large amounts of antisense rRNA probes). the Ambion™ RPA II™ and RPA III™, and HybSpeed™ RPA Kits are also available for quick and accurate analysis of RNA samples.
For accurate RT-PCR analysis, we provide the Ambion™ QuantumRNA™ family of kits (with primer and Competimer™ sets for 18S rRNA and ß-actin) for relative quantitation, and the Competitive Quantitative RT-PCR Kits (pre-made Armored RNA™ Competitors for human interleukin RNAs) for absolute quantitation. We also provide an RT-PCR Competitor Construction Kit for synthesizing RNase-resistant RNA competitors for RT-PCR.