Search
LanthaScreen Cellular Assays

- Screen quickly with a simple and homogenous high-throughput screening (HTS) – amenable assay format
- Interrogate the effects of your compounds on specific signaling events within a pathway
- Analyze multiple post-translational modifications using physiological pathway kinetics
To view the available LanthaScreen cellular assays, plus extensive validation data and protocols specific to each post-translational modification available, please see our LanthaScreen cellular assay validation table.
Assay Principle
The cellular response to environmental cues is mediated through a complex network of signal transduction pathways. Alteration or disruption of components within these pathways often contributes to various diseases, including cancer and metabolic and inflammatory disorders. The diversity of validated drug targets in these signal transduction cascades provides both challenges and opportunities in drug discovery. Intermediate signaling events, such as phosphorylation and ubiquitination, are bona fide targets for assay development. Until now, there have been a limited number of HTS-compatible methods available to interrogate, in a cell-based format, the regulated activity of enzymes responsible for posttranslational modifications.
Combining the simplicity of biochemical assays with the intricacy of cellular systems, Invitrogen has developed time-resolved fluorescence resonance energy transfer (TR-FRET)-based assays to monitor protein modification within the native cell signaling environment. By expressing the substrate of interest as a fusion with green fluorescent protein (GFP), modification-specific antibodies labeled with the TR-FRET donor partner terbium (Tb) can be used to quantitatively monitor these modifications following cell lysis (Figure 1).
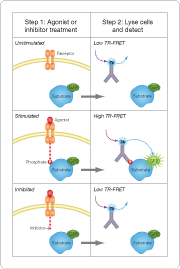
Figure 1. Schematic of LanthaScreen cellular assay. The principle and methods for this assay format are simple. 1) Phosphorylation of the kinase substrate expressed as a GFP-fusion protein is either stimulated with an agonist to activate the pathway or treated with an inhibitor to interrupt the signaling event. No protein modification occurs in unstimulated cells. 2) Cells are lysed in a buffer that contains a Tb-labeled phosphorylation site-specific antibody. The detection of the protein modification event is measured on a TR-FRET-compatible plate reader. Little or no TR-FRET is observed with unstimulated or inhibited cells whereas stimulated cell samples display high TR-FRET.
Optimize workflows for HTS applications
Compared to classical and established methods for phosphoprotein analysis, such as western blotting and ELISA, the workflow for a LanthaScreen cellular assay is quick, designed for HTS screening in a 384-well plate, and highly amenable to automation (Figure 2).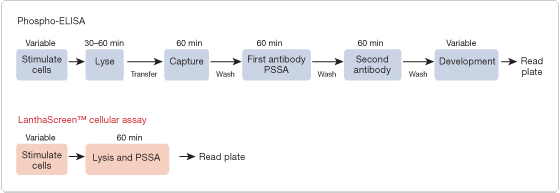
Figure 2. Comparison of LanthaScreen cellular assay and phosphoELISA workflow. The LanthaScreen cellular assay platform offers a high-throughput alternative for compound screening and profiling. In contrast to the traditional ELISA-based approach for phospho-protein analysis, this assay format requires minimal hands-on time and can be run in as few as 2−3 hours as opposed to the typical 6- to 8-hour time commitment with classical phosphoELISA™ assays. (Phosphorylation site-specific antibodies; PSSA).
This assay format has no wash steps, and the assays can be run following an “addition only” protocol, requiring little hands-on time and eliminating the need for extra manual touches and liquid transfer steps. Moreover, the data obtained in the LanthaScreen cellular assay format are equivalent to those generated using a phosphoELISA (Figure 3).
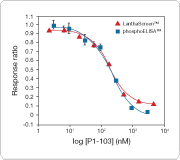
Figure 3. Equivalent experimental data generated using a LanthaScreen cellular assay and a phosphoELISA. HEK293E cells were treated with different concentrations of the dual PI3K/mTOR inhibitor PI-103 and subsequently analyzed using the PRAS40 [pT246] phosphoELISA kit from Invitrogen. The inhibitor data are overlaid with the results from the LanthaScreen PRAS40 [T246] HEK293E assay, with nearly identical IC50 values (~200 nM). Together, these data indicate that the LanthaScreen cellular assay format is a suitable surrogate for established methods for phosphoprotein analysis.
Support
Technical resources
Posters
Publications
For Research Use Only. Not for use in diagnostic procedures.