Search
Overview of Cell Lysis and Protein Extraction
Structure and diversity of cells
All cells have a plasma membrane, a protein-lipid bilayer that forms a barrier separating cell contents from the extracellular environment. Lipids comprising the plasma membrane are amphipathic, having hydrophilic and hydrophobic moieties that associate spontaneously to form a closed bimolecular sheet. Membrane proteins are embedded in the lipid bilayer, held in place by one or more domains spanning the hydrophobic core. In addition, peripheral proteins bind the inner or outer surface of the bilayer through interactions with integral membrane proteins or with polar lipid head groups. The nature of the lipid and protein content varies with cell type and species of organism.
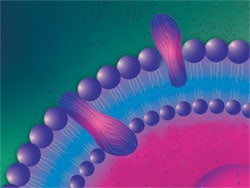
Cell membrane structure. Illustration of a lipid bilayer comprising outer plasma membrane of a cell.
In animal cells, the plasma membrane is the only barrier separating cell contents from the environment, but in plants and bacteria the plasma membrane is also surrounded by a rigid cell wall. Bacterial cell walls are composed of peptidoglycan. Yeast cell walls are composed of two layers of ß-glucan, the inner layer being insoluble to alkaline conditions. Both of these are surrounded by an outer glycoprotein layer rich in the carbohydrate mannan. Plant cell walls consist of multiple layers of cellulose. These types of extracellular barriers confer shape and rigidity to the cells. Plant cell walls are particularly strong, making them very difficult to disrupt mechanically or chemically. Until recently, efficient lysis of yeast cells required mechanical disruption using glass beads, whereas bacterial cell walls are the easiest to break compared to these other cell types. The lack of an extracellular wall in animal cells makes them relatively easy to lyse.
There is no universal protocol for protein sample preparation. Sample preparation protocols must take into account several factors, such as the source of the specimen or sample type, chemical and structural heterogeneity of proteins, the cellular or subcellular location of the protein of interest, the required protein yield (which is dependent on the downstream applications), and the proposed downstream applications. For instance, bodily fluids such as urine or plasma are already more or less homogeneous protein solutions with low enzymatic activity, and only minor manipulation is required to obtain proteins from these samples. Tissue samples, however, require extensive manipulation to break up tissue architecture, control enzymatic activity, and solubilize proteins.
The quality or physical form of the isolated protein is also an important consideration when extracting proteins for certain downstream applications. For instance, applications such as functional enzyme-linked immunosorbent assay (ELISA) or crystallography require not only intact proteins but also proteins that are functionally active or retain their 3D structure.

Examples of protein sources for sample collection. Proteins can come from many sources, including the following: native sources such as mammalian cell cultures, tissues or bodily fluids; overexpression in a model system such as bacteria, yeast, insect or mammalian cells; monoclonal antibodies from hybridoma cells; or plant cells used in agricultural biotechnology.
Cell lysis and protein extraction
Historically, physical lysis was the method of choice for cell disruption and extraction of cellular contents; however, it often requires expensive, cumbersome equipment and involves protocols that can be difficult to repeat due to variability in the apparatus (such as loose-fitting compared with tight-fitting homogenization pestles). Also, traditional physical disruption methods are not conducive for high-throughput and smaller volumes typical of modern laboratory research.

Cell or tissue disruption methods. Many cell lysis methods have been developed to obtain the best possible yield and purity for different species of organisms, sample types (cells or tissue), and target molecule or subcellular structure. Cell lysis methods can be divided into two main categories, reagent-based and physical disruption.
In recent years, detergent-based lysis methods have become the norm. Through empirical testing by trial and error, different detergent-based solutions composed of particular types and concentrations of detergents, buffers, salts and reducing agents have been developed to provide the best possible results for particular species and types of cells. Detergents have both lysing and solubilizing effects.

Isolation of high quality protein samples is the cornerstone to completing successful downstream applications.
By careful optimization of physical disruption techniques, detergent–buffer solutions and density gradient methods, procedures have been developed to enable separation of subcellular structures or classes of compounds. For example, with the appropriate detergents, hydrophobic membrane proteins can be solubilized and separated from hydrophilic proteins. A combination of tools and steps enable intact nuclei, mitochondria and other organelles to be isolated for study or protein solubilization.

Nucleus, mitochondria and other organelles inside a cell.
Cell lysis disturbs the carefully controlled cellular environment, allowing endogenous proteases and phosphatases to become unregulated. As a result, extracted proteins become degraded or artifactually modified by the activities of these molecules.
To prevent these effects and obtain the best possible protein yield in cell lysis, protease and phosphatase inhibitors are added to the lysis reagents. Numerous compounds have been identified and used to inactivate or block the activities of proteases and phosphatases by reversibly or irreversibly binding to them.

Protein phosphorylation is preserved in cell and tissue extracts. Relative levels of total and phosphorylated protein from extracts prepared in the absence or presence of phosphatase inhibitors were determined by western blot analysis. The degree of protein inhibition for acid and alkaline phosphatase activity was determined in mouse brain extract after treatment with Thermo Scientific Pierce Phosphatase Inhibitor Tablets or another commercially available phosphatase inhibitor tablet. Percent inhibition is indicated.
When the goal of cell lysis is to purify or test the function of a particular protein, special attention must be given to the effects of the lysis reagents on the stability and function of the protein(s) of interest. Certain detergents will inactivate the function of particular enzymes, and long-term stability of extracted/purified proteins often requires that they be removed from the initial lysis reagents and/or stabilized by addition of particular compounds.
Select products
Some cell lysis and protein solubilization methods cause the denaturation of proteins. Functional tests with such proteins require that they be renatured. Many proteins spontaneously refold into their native, functional structures when the denaturing solubilization reagents are removed by dialysis. Other proteins, however, will fold into non-functional and even insoluble forms by this process. In such cases, specialized sets of buffer conditions must be tested to identify those that promote the highest possible yield of properly refolded protein.
| Buffer | Guanidine | L-Arginine | Redox environment | Protein precipitation | Relative percent recovery |
|---|---|---|---|---|---|
| 1 | 0.4M | 0M | 5mM DTT | YES (high) | 2% |
| 2 | 0.4M | 0.4M | 2mM GSH 0.2mM GSSG | YES (mid) | 34% |
| 3 | 0.4M | 0.8M | 2mM GSH 0.4mM GSSG | NO | 81% |
| 4 | 0.9M | 0M | 2mM GSH 0.2mM GSSG | YES (low) | 47% |
| 5 | 0.9M | 0.4M | 2mM GSH 0.4mM GSSG | NO | 97% |
| 6 | 0.9M | 0.8M | 5mM DTT | NO | 7% |
| 7 | 1.4M | 0M | 2mM GSH 0.4mM GSSG | NO | 100% |
| 8 | 1.4M | 0.4M | 5mM DTT | NO | 7% |
| 9 | 1.4M | 0.8M | 2mM GSH 0.2mM GSSG | NO | 7% |
Results for refolding lysozyme using the Pierce Protein Refolding Kit. Each buffer contains the indicated denaturant and redox concentrations as well as 50 mM Tris, 18 mM NaCl, 8 mM KCl, 1 mM EDTA; pH 8.2. Recovery is reported as a percentage of the trial (Buffer 7) having highest activity after refolding. Compared to non-denatured controls, Trial 7 represents greater than 90% of the solubilized lysozyme refolding. (Polyethylene Glycol (PEG); Dithiothreitol (DTT); Reduced glutathione (GSH); Oxidized glutathione (GSSG)).
Protein Preparation Handbook
The Protein Preparation Handbook provides useful information on our broad portfolio of reagents and tools for protein extraction, clean-up, immunoprecipitation and purification. Practical information, selection guides, and relevant data are included to help you improve your protein yield and downstream analysis.
Recommended reading
- Walker JM (2009) The Protein Protocols Handbook. Third Edition. New York (NY): Springer-Verlag New York, LLC.
- Vallejo FL, Rinas U (2004) Strategies for recovery of active proteins through refolding of bacterial inclusion body proteins. Microb Cell Fact 2:3(1):11.
For Research Use Only. Not for use in diagnostic procedures.
