Search
Organelle Dynamics in Live Cells
Examining cellular pathways in both their spatial and temporal context is critical for understanding cell development and functioning. Molecular Probes CellLight reagents provide a simple and effective method for introducing targeted intracellular labels into live cells, enabling unambiguous visualization of organelles and other cellular structures in live mammalian cells by fluorescence microscopy (Figure 1).The secretory pathway involves a series of complex and highly regulated steps that ultimately leads to the delivery of proteins out of the cell. The endoplasmic reticulum (ER) and Golgi apparatus are the key membrane-bound intracellular compartments involved in the processing and posttranslational modification of proteins prior to their delivery. These compartments exist in a dynamic equilibrium mediated by anterograde traffic from the ER to the Golgi, and reciprocal retrograde trafficking from the Golgi to the ER. This transport is mediated by small GTPases, which participate in the budding of transport vesicles. Similar processes govern lysosome-mediated organelle degradation (autophagy). When CellLight reagents are combined with Molecular Probes classic organelle stains, these dynamic organelle processes can be studied in their native cellular context (Figures 2 and 3).
CellLight reagents are provided in a ready-to-use format—simply add, incubate, and visualize. These reagents open up new avenues for multiparametric study of dynamic cellular events in live-cell context.
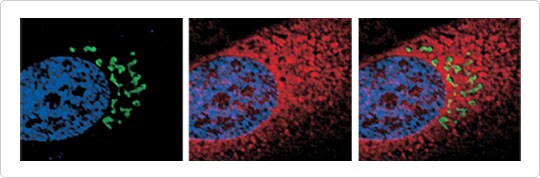
Figure 1. Labeling of nuclei, Golgi apparatus, and ER in live HeLa cells. Nuclei were labeled with Hoechst 33342 (blue), the Golgi apparatus was labeled with CellLight Golgi-GFP (green), and the ER was labeled with CellLight ER-RFP (red). The close relationship between the ER and the Golgi apparatus is evident in this high-resolution image, with the perinuclear Golgi apparatus appearing in “pockets” of the ER. Imaging was performed on a DeltaVision Core microscope.
Lysosome-mediated Organelle Degradation
 | Figure 2. Lysosome-mediated organelle degradation (autophagy) visualized with CellLight™ and LysoTracker reagents. Cells were transduced with CellLight Mito-GFP (green) and subsequently serum-starved for 24 hr to induce autophagy. Nuclei were labeled with Hoechst 33342 (blue) and total lysosomal mass was assessed with LysoTracker Red DND-99 (red). Under control conditions (Top), a normal population of numerous small lysosomes was observed. Under serum starvation (Bottom), a smaller number of large lysosomal structures were observed in areas of the cell devoid of mitochondria. Imaging was performed on a DeltaVision Core microscope. |
ER-to-Golgi Trafficking
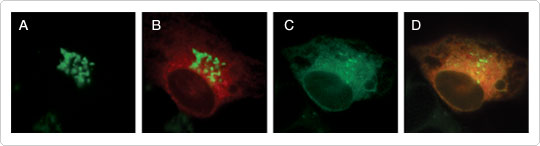
Figure 3. ER-to-Golgi trafficking visualized with CellLight reagents.The ER and Golgi apparatus were labeled with CellLight ER-RFP (red) and CellLight Golgi-GFP (green) and imaged before (A, B) and following (C, D) brefeldin A treatment. Brefeldin A blocked anterograde transport from the ER to the Golgi apparatus, leaving only retrograde transport intact. As a result, the Golgi stacks redistributed to the ER via unperturbed retrograde trafficking, colocalizing with the ER-targeted RFP and appearing as orange fluorescence. To resolve the dispersed Golgi GFP signal in the ER after brefeldin A treatment, the images in (C) and (D) were adjusted. Identical adjustments in gain made to (A) and (B) revealed no diffusely localized Golgi GFP prior to brefeldin A treatment (data not shown). Images were acquired every 30 sec for 30 min following treatment. Imaging was performed on a DeltaVision Core microscope.
NOTE: Organelle Lights reagents were the original products used for these studies. However, subsequent to this articles' publication in March, 2010, we have released a new and improved, next generation version of Bacmam reagents, including CellLight reagents. We have found that these new products perform equal to and in most cases, better than, the previous versions. Please don't hesitate to contact our technical support team if you have any questions or concerns.
Mitochondrial Biology Poster
Probes for Mitochondrial Morphology and Function
Download This Article
Get a copy of this article as it appears in the print version of BioProbes 62.
Learn More
Find more information on organelle-reagents.
Quick Product View
See a complete listing of the products discussed in this article.
View products
For Research Use Only. Not for use in diagnostic procedures.
