Search
Accuracy and Precision with the Countess II FL Automated Cell Counter
The four pillars of achieving accurate and precise cell counts
Many biological science applications require an accurate, precise measurement of the number of cells and their viability prior to downstream analysis. This assessment can be done manually with a traditional hemocytometer and microscope or with an automated cell counter, such as the Invitrogen Countess II FL Cell Counter.
When manually counting cells using a glass hemocytometer together with a microscope, count-to-count variability of a single sample by an experienced cell biologist is commonly 10% or more. When comparing variation of single-sample counts across multiple scientists, counting variability commonly exceeds 20%. Automated cell counters minimize the subjective nature inherent in manual counting as well as user-to-user differences in total cell count and viability while helping to save time. Users of Countess II Automated Cell Counters should typically see count-to-count variability <5% CV% when following the tips outlined in this paper.
Regardless of the chosen cell counting platform, achieving an accurate and precise count from repetitive counts of a single sample or across multiple samples is critical, yet sometimes challenging. By remembering the four pillars of accurate and precise cell counting listed below and ranked in order of importance, one can minimize variability and maximize cell counting confidence.
Sample preparation
How the cells of interest are prepared can have a dramatic effect on cell counts for manual and automated counting. Maintaining uniformity between samples is vital to being able to determine if differences are caused by an experimental treatment or by the sample preparation conditions.
Here are a few sample preparation tips that will increase your counting accuracy and precision.
- Standardize cell handling steps. In many cases, cell viability and concentration comparisons are made across multiple samples, which may come from different flasks or culture plates. If so, it is important to realize that slight differences in cell handling steps including proteolysis, pipetting, or centrifugation procedures can have a dramatic effect on the observed total cell count and viability reported from sample to sample.
- Ensure cell suspension is homogenously mixed. A homogenous suspension can be achieved by using a brief, gentle vortexing motion or by manually agitating the tube by hand—some refer to this the “finger flick” technique. Allowing your cells to sit on the benchtop for a minute prior to pipetting will result in the settling of cells toward the bottom of the tube, creating an artificial concentration gradient and a potential source of error. The longer a cell sample sits, the greater the concentration gradient will become, thereby increasing variability. To minimize pipetting variability, mix the sample as noted above, then pipette from the middle of the tube rather than the bottom where there may be an artificial concentration of cells.
- Minimize debris whenever possible. Regardless of the chosen counting or staining method, having a sample with significant amounts of debris complicates counting and viability assessment. The amount of debris varies based on cell type, culture conditions, preparation protocol, and the reagents being used. Taking into consideration the possible causes of debris and taking steps to minimize them, when possible, will improve the accuracy of cell counting and viability results.
Common tips to minimize debris in cell samples:- Centrifuge trypan blue before use
- Use fresh staining reagents (trypan blue, NucBlue Live, propidium iodide, etc.)
- Use sterile-filtered media and buffers
- Passage cells at required intervals
Setting focus correctly
Obtaining the correct focus is critical for obtaining accurate viability information. Focus consistency between cell counts is critical to minimize sample-to-sample variance. When manually counting cells, setting focus is subjective and can vary between users. However, many of the latest automated image-based cell counters utilize an auto-focus feature that is not only convenient, but also eliminates the subjective adjustment of focus. This alone will enable better counting consistency between samples.
The example images in Figure 1 reflect identical fields of view (FOV) of the same sample and illustrate the difference that can be seen in both manual and automated counting methods when focus is not set correctly. The image in Figure 1A demonstrates correct focus, while the focus in Figure 1B is incorrect. The incorrectly focused image is falsely identifying live cells as dead cells. Remember, regardless of your counting method, cells can only be counted if they are visible within the field of view being counted.
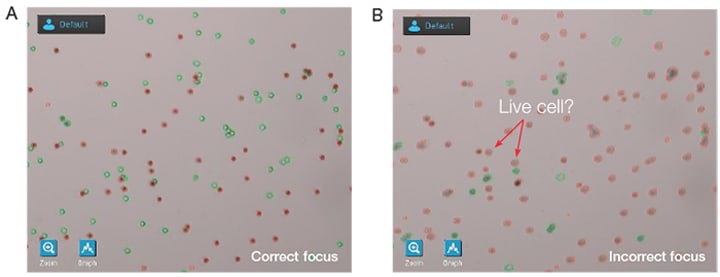

Figure 1. (A) The live cells (circled in green) have bright centers and slightly darker boarders when the focus is set properly. (B) Dead cells (circled in red) may look slightly out of focus, but will present a uniform dark staining pattern if trypan blue is present.
Staining and light intensity
Bright-field counting
Trypan blue solution, 0.4%, is routinely used to rapidly assess cell viability. The dye exclusion test is based upon the concept that viable cells do not take up impermeable dyes (like trypan blue), but dead cells are permeable and take up the dye, which produces results as illustrated in Figure 1A. Unlike manual counting methods where users must not only manually adjust focus but also bright-field light intensity between samples, automated cell counters like the Countess II FL are able to automatically adjust lighting and focus to obtain optimal image quality used to determine viability. If the light intensity is too high the screen will appear washed out (Figure 2A). If the light intensity is too low the background will be too dark (Figure 2B). Light intensity settings that are too high or too low will result in an inaccurate cell count.
Fluorescent detection
When using an automated cell counting instrument that is able to detect fluorescence there are two key variables that should be considered, signal-to-noise ratio and background. It is important in any fluorescent staining study to minimize background (e.g., noise) while maximizing signal, as doing so will increase sensitivity while maximizing signal-to-noise ratio. One requirement to achieve this goal is to correctly set the excitation intensity by adjusting the light source. See Figure 3 for a comparison of good excitation intensity to excitation intensity that is too high.
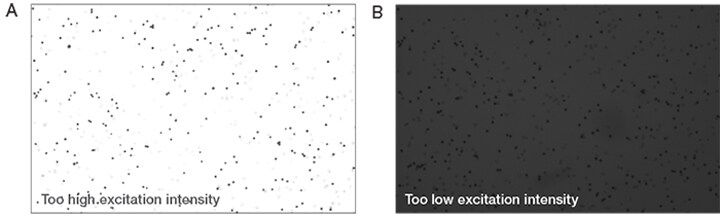
Figure 2. (A) The bright-field light intensity is set too high. (B) The light intensity is too low.
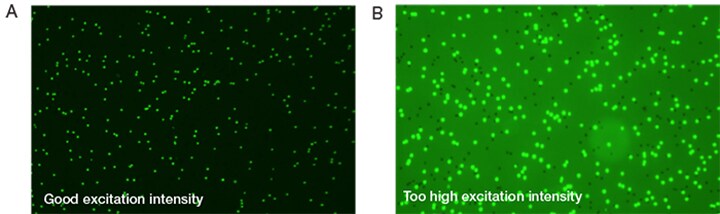
Figure 3. (A) The optimal fluorescent light intensity results in a good image. (B) Reflects an image with the light intensity set too high.
Gating
Finally, when using automated cell counters, the ability to include or exclude (gate) cells from a count is easily achievable compared to manual counting, yet also a source of inconsistent counts if different gating strategies are used between samples. As a general rule of thumb, similar gating strategies should be used when comparing samples. If using the Countess II or II FL, adjust the gating parameters via the “More” button on the results screen and save the gating settings as a “Profile” (Figure 4) for easy recall as needed.
Gating parameters commonly used alone or in combination include: cell size, brightness, circularity, and fluorescence intensity. Together, these parameters allow a user to fine-tune precisely what is included or excluded for a given cell counting application.
Circularity
Debris can be present in dramatically different amounts depending upon the cell type, culture conditions, preparation protocol, and the reagents being used. Some classic debris examples include particulate from contaminated trypan blue, and cell debris from dead or dying cells in old cultures or whole blood preparations. In most cases, the debris is irregular in shape and stains dark with trypan blue, which allows it to be efficiently ‘gated out’ using the circularity parameter in many automated cell counters, including Countess II automated cell counters.
In Figure 5A and 5B, a sample containing a significant amount of debris was counted in bright-field mode without and with gating applied, respectively. Some of the debris was detected as dead cells; since the debris noted was small in size, irregular in shape, and relatively faint compared to background, a gating strategy was employed via the “More” button. The image on the results screen is updated in real time for customer feedback for visual confirmation.
Size
Bright-field gating by size is commonly done in applications where multiple cell types could be present (e.g., neuronal and stem cell cultures, blood samples, or when debris is present). Using automated cell counters, one can easily set gates to include or omit specific cells from being counted (Figure 6). With the Countess II automated cell counters, it is important to note that gating can be independently performed on both live and dead cell populations, thus allowing for greater flexibility. Furthermore, these gating settings can be saved for future use.
Brightness
Gating based on fluorescence increases gating strategy flexibility. In many cases, users simply want to know how many cells are dim vs. bright or expressing a fluorescent protein. Figure 7A shows a GFP-expressing sample with two populations, dim and bright; the dim cells have been gated out revealing that 13% of the cells counted are strongly expressing GFP. In b, two fluorescence colors are present, both of which can be independently gated on the basis of size, brightness, circularity, and fluorescent intensity via the “More” button present on the Results screen. Countess II FL can save these parameters for consistency between samples and future use.

Figure 4. Adjustments can be made for each threshold parameter and then saved as needed to create the requisite profile.
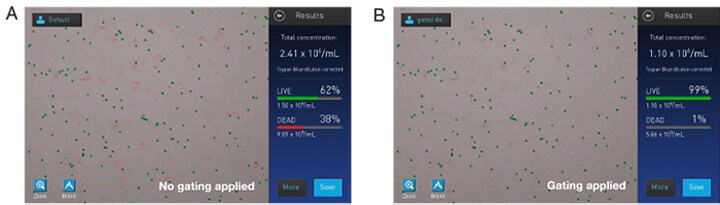
Figure 5. (A) No gating applied; note faint debris associated with red circles. (B) Size, brightness, and circularity gates applied; note lack of debris circled.

Figure 6. The light grey bars indicate that cells less than 10 μm have been excluded from the indicated cell count results.
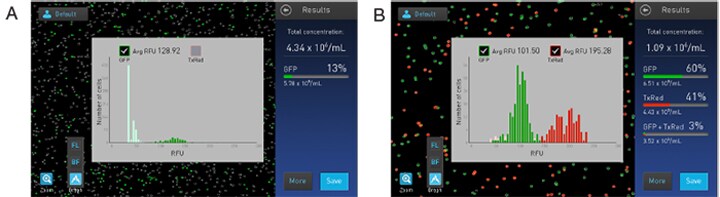
Figure 7. (A) The light green bars around 50 RFUs have been gated out of the GFP-positive count. (B) Two fluorescent colors are present, each of which can be gated by size, brightness, circularity, and fluorescent intensity.
Summary
Variability in cell counting results can have a variety of causes. Sample preparation, focus settings, light intensity settings, and gating strategies can all have an impact on results. Use of an automated cell counter that applies autofocus, auto bright-field lighting, and has multiple gating settings will improve the reliability of cell counting and viability reporting. The ability to save profiles for consistent application of settings across samples can help improve result accuracy.
Resources
Publications
Application notes
Technical resources
Instrument support & demo
For Research Use Only. Not for use in diagnostic procedures.
For Research Use Only. Not for use in diagnostic procedures.