Search
How TaqMan Assays Work

Applied Biosystems™ TaqMan® real-time PCR assays consist of target-specific primers and one or more probes optimized for specific types of measurements. The mechanisms of action of different types of assays are described below.
TaqMan® Gene Expression Assays are based on 5' nuclease chemistry, which uses a fluorogenic probe to enable the detection of a specific PCR product as it accumulates during PCR.
Each assay contains:
- A pair of unlabeled primers
- A TaqMan probe with a FAM™ or VIC™ dye label on the 5' end, and a minor groove binder (MGB) and nonfluorescent quencher (NFQ) on the 3' end
The figure below depicts the TaqMan Gene Expression Assay process.

- At the start of real-time PCR, the temperature is raised to denature the double-stranded cDNA. During this step, the signal from the fluorescent dye on the 5' end of the TaqMan probe is quenched by the NFQ on the 3' end.
- In the next step, the reaction temperature is lowered to allow the primers and probe to anneal to their specific target sequences.
- Taq DNA polymerase synthesizes new strands using the unlabeled primers and the template. When the polymerase reaches a TaqMan probe, its endogenous 5' nuclease activity cleaves the probe, separating the dye from the quencher.
With each cycle of PCR, more dye molecules are released, resulting in an increase in fluorescence intensity proportional to the amount of amplicon synthesized.
TaqMan MGB probes contain a minor groove binding moiety that enhances the Tm differential between matched and mismatched probes. In addition, TaqMan MGB probes contain a nonfluorescent quencher that enhances spectral resolution when using multiple dyes in a reaction.
Taq Talk Episode 4 — How TaqMan real-time PCR assays work
TaqMan genotyping assays (TaqMan® SNP Genotyping Assays and TaqMan® Drug Metabolism Genotyping Assays) consist of pre-optimized PCR primer pairs and two probes for allelic discrimination .
Each assay contains:
- A pair of unlabeled primers
- Two TaqMan probes (one with a FAM dye label and one with a VIC dye label) on the 5’ end and minor groove binders (MGB) and nonfluorescent quenchers (NFQ) on the 3´ end.
TaqMan genotyping assays are used to amplify and detect specific alleles in genomic DNA (gDNA). The figure below depicts the TaqMan SNP Genotyping Assay process.
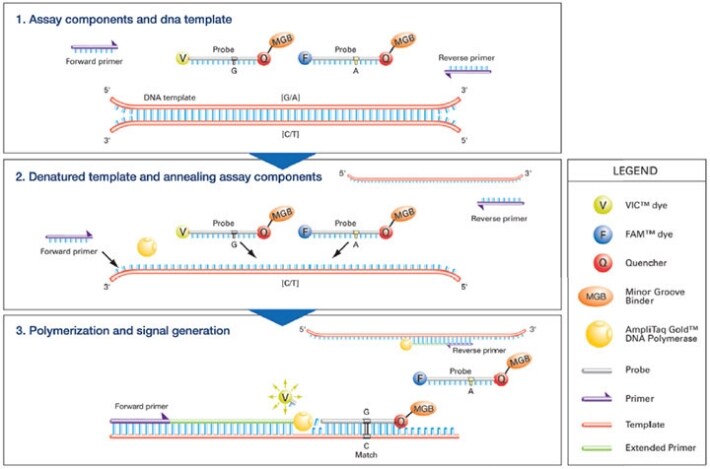
- Genomic DNA is introduced into a reaction mixture consisting of TaqMan® Genotyping Master Mix, forward and reverse primers and two TaqMan® MGB Probes.
- Each TaqMan MGB Probe anneals specifically to a complementary sequence, if present, between the forward and reverse primer sites. When the probe is intact, the proximity of the quencher dye to the reporter dye suppresses the reporter fluorescence.
- The exonuclease activity of AmpliTaq Gold® DNA Polymerase cleaves only probes hybridized to the target. Cleavage separates the reporter dye from the quencher dye, increasing fluorescence by the reporter. The increase in fluorescence occurs only if the amplified target sequence is complementary to the probe. Thus, the fluorescence signal generated by PCR amplification indicates which alleles are in the sample.
TaqMan SNP Genotyping Assays › TaqMan Drug Metabolism Genotyping Assays ›
TaqMan® Copy Number Assays are run together with a TaqMan® Copy Number Reference Assay in a duplex real-time Polymerase Chain Reaction (PCR). The Copy Number Assay detects the target gene or genomic sequence of interest and the Reference Assay detects a sequence that is known to be present in two copies in a diploid genome.

During PCR:
a. A TaqMan® Copy Number Assay, a TaqMan® Copy Number Reference Assay, TaqMan® Genotyping Master Mix, and a gDNA sample are mixed together in a single well or tube.
b. The gDNA template is denatured and each set of assay primers anneals to its specific target sequences. Each TaqMan® probe anneals specifically to its complementary sequence between forward and reverse primer binding sites. When each oligonucleotide probe is intact, the proximity of the quencher dye to the reporter dye causes the reporter dye signal to be quenched.
c. During each round of PCR, the target and reference sequences are simultaneously amplified by AmpliTaq® Gold DNA Polymerase. This enzyme has a 5′ nuclease activity that cleaves probes that are hybridized to each amplicon sequence. When an oligonucleotide probe is cleaved by the AmpliTaq Gold DNA Polymerase 5′ nuclease activity, the quencher is separated from the reporter dye increasing the fluorescence of the reporter. Accumulation of PCR products can be detected in real time by monitoring the increase in fluorescence of each reporter dye at each PCR cycle.
This method of relative quantitation is used to determine the relative copy number of the target of interest in a gDNA sample, normalized to the known copy number of the reference sequence.
Ask TaqMan #34 Copy Number Variation – How Does It Work?
Quantification using TaqMan® Advanced miRNA Assays requires a cDNA synthesis step followed by the detection using real-time PCR.

The TaqMan Advanced miRNA Assay workflow.
(A) The cDNA synthesis workflow incorporates a 3´ poly(A) addition and 5´ adapter ligation step, followed by reverse transcription with a universal RT primer. An optional miR-Amp reaction is then performed to uniformly increase miRNA cDNA prior to detection by qPCR.
(B) Each TaqMan Advanced miRNA Assay employs a TaqMan minor groove binder (MGB) probe that specifically anneals to a complementary sequence between the forward and reverse primer sites. The MGB modification increases melting temperature without increasing probe length, allowing for design of shorter probes. When the probe is intact, the proximity of the reporter dye to the nonfluorescent quencher (NFQ) results in suppression of reporter fluorescence. Probe cleavage by DNA polymerase during primer extension separates the reporter dye from the NFQ, resulting in increased fluorescence of the reporter.
Ask TaqMan #31 Finding The Right MicroRNA Assays
TaqMan® Mutation Detection Assays are based on a novel competitive allele-specific TaqMan (castPCR™) technology, which combines allele-specific TaqMan qPCR with an allele-specific MGB blocker oligonucleotide to effectively suppress nonspecific amplification of the nontarget allele.

A mutant allele assay or wild type allele assay is composed of an allele-specific primer (red), locus-specific primer (purple), locus-specific TaqMan probe (green), and allele-specific blocker (blue). The level of mutation present in the sample is calculated based on the ΔCt for a mutant allele assay and the corresponding wild type allele assay, or for a mutant allele assay and a gene reference assay.
TaqMan® Protein Assays are an adapted form of proximity ligation assays (PLA™), which combine antibody–protein binding with detection of the reporter nucleic acid by real-time PCR.
TaqMan Protein Assay Workflow

| Lyse samples (cells or tissues) using a gentle 1-step procedure in a buffered non-ionic detergent, with no further sample clean-up or purification steps required. |
|---|---|
Binding | The assay probes are target-specific antibodies conjugated to oligonucleotides through a biotin–streptavidin linkage. Each oligonucleotide in the pair presents a 5′ or 3′ end; these are brought into proximity when the antibodies on the assay probes bind to two different epitopes on the target protein. |
Ligation/inactivation | The substrate for ligase is a bridge structure formed by hybridization of a third oligonucleotide to the oligonucleotide ends of the assay probe pair. This structure forms preferentially when the assay probes are brought into proximity by binding to the target protein. |
TaqMan Fast real-time PCR | The ligation product serves as a DNA template for real-time PCR using a TaqMan Protein Assay. The mechanism of action is the same as that of the gene expression assay, using unlabeled primers and a TaqMan probe with a fluorescent label on the 5´ end and a nonfluorescent quencher (NFQ) on the 3´ end. |
Data analysis | Analyze TaqMan Protein Assay data using our free ProteinAssist™ Software package. |
For Research Use Only. Not for use in diagnostic procedures.