Search
Luciferase Reporters
Introduction to bioluminescence
Light emission has been used to detect experimental changes in biological assays for almost 100 years (1), and while the list of applications that use emitted light as a detection strategy is long, the type of light emitted varies. Luminescence is light emission as a result of a chemical reaction without the production of heat or any thermal changes. This type of light is clearly different from incandescence, which generates heat and is the reason why incandescent light bulbs get hot during use. Luminescence can be separated into two types:
- Bioluminescence – light emitted from a biological source
- Chemiluminescence – light emitted from a non-biological source due to a chemical reaction

Application Note-Luciferase Assay and Bio Tek Synergy Microplate Readers
This application note describes validation and comparison of single-step flash and dual-spectral luciferase reporter gene assays using the Synergy line of microplate readers.
Select products
Bioluminescence vs. fluorescence
Fluorescence is another type of emitted light commonly used in biological research and is the product of a fluorophore, a molecule that absorbs the energy from a light source and then emits light at a different wavelength. Bioluminescence, on the other hand, differs from fluorescence in that the excitation energy is supplied by an enzymatic reaction rather than from a source of light. While both bioluminescence and fluorescence are widely used in scientific applications, bioluminescent reporters display an ultrasensitive detection capacity and have a wider dynamic range compared to fluorescent reporters because of the enzymatic nature of the bioluminescent reporter.
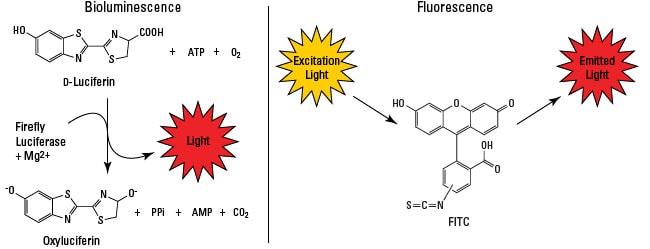
Bioluminescence vs. fluorescence. Bioluminescence (left) is emitted from the reaction of luciferase enzyme and its substrate, such as firefly luciferase and luciferin, respectively. Cofactor requirements (e.g., ATP, O2) vary depending on the luciferase used. Fluorescence (right) is the product of a fluorophore (e.g., FITC, Alexa Fluor dyes) absorbing the energy from a light source and emitting the light energy at a different wavelength.
Fluorescent reporters are susceptible to photo bleaching, provide low quantum yields and have greater protein stability in cell-based assays compared to bioluminescent reporters, which make them less amenable for use as real-time reporters. Cellular components also have auto fluorescent properties, which increases the non-specific background and decreases the sensitivity of fluorescent detection in cell-based assays. Conversely, cellular components have no inherent bioluminescence, allowing for greater sensitivity with bioluminescent assays.
However, fluorescent reporters are more useful for visualizing targets in live cells than bioluminescent proteins, because fluorophores do not require cofactors or exogenous substrates for activity and are more stable than bioluminescent reporters.
Select products
Bioluminescence in nature
Uses of bioluminescence in nature are quite diverse and are exhibited by both unicellular organisms like bacteria and dinoflagellates and higher-order organisms such as fish and insects. Bioluminescence is used by both terrestrial and aquatic organisms, although it is more commonly seen in marine animals, especially those living in the extreme depths of the ocean. Bioluminescence serves many natural purposes for these organisms.
Defense: A strong bioluminescent signal can be used by prey to temporally blind an attacking predator, giving the prey a chance to escape. The light produced by the prey also puts the potential predator at the risk of being detected by other predators.
Camouflage: Animals that normally dwell in oceanic dark zones, including firefly squids, krill, dogfish sharks and hatchet- and lantern fish, come to the surface to feed at night. These animals use counter-illumination to blend into surrounding areas illuminated by moonlight. Similar camouflage is also used by certain species of shallow-water squid that give off light to blend in with the moonlight.
Feeding: Predators often use bioluminescence to attract prey. The anglerfish, for example, lures prey using an elongated dorsal spine that supports a light-producing organ that houses bioluminescent bacteria. Also, some octopi, including Stauroteuthis syrtensis, have suckers that are lined with organs that produce blue-green light that are also used to attract prey.
Mating: Terrestrial animals such as fireflies, glow worms and marine animals like the octopod Japetella diaphana use bioluminescence to attract a mate.
Bioluminescent reaction
Two agents are essential for a bioluminescent reaction to occur: luciferase, the enzyme that catalyzes the reaction, and the luciferase substrate. A wide variety of luciferase enzymes have been discovered, although the general mechanism of the light-producing reaction for all of the enzymes is the oxidative decarboxylation of the luciferase substrate in the presence of O2 to yield photon emission (light).
Luciferases likely evolved at multiple stages during the course of history to yield enzymes that differ in expression pattern, substrate specificity, cofactor requirement and enzyme kinetics. The result of these evolutionary changes is that some organisms, such as bacteria and fungi, can emit light continuously, while other organisms emit bioluminescent flashes with varying duration and intensity. For example, the luminescence from a dinoflagellate is short, lasting 0.1 seconds, while that from jellyfish can last tens of seconds.
The variation in bioluminescent output by different luciferase enzymes has allowed scientists to classify them based on their output kinetics to accommodate different experimental designs. Luciferase enzymes with flash kinetics have maximum sensitivity because of high signal intensity, although the emitted light also rapidly decays. Conversely, enzymes with glow kinetics are less sensitive but stably emit light for at least 60 minutes.
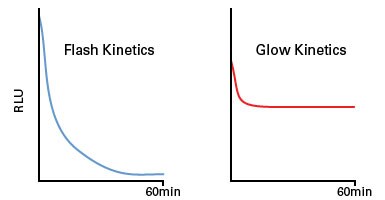
Luciferase flash vs. glow kinetics. Schematic representation of the bioluminescent output of luciferase enzymes with flash and glow kinetics as a function of time. RLU: relative light units.
Select products
Luciferase enzymes
Luciferase enzymes are active in their native state and lose activity upon unfolding or denaturation, a property that has been utilized to study factors that influence proper protein folding. Also, luciferases such as Gaussia, Metridia and Cypridina are naturally secreted and therefore have been used to study the mechanisms and regulation of the secretory pathway.
| Organism | Luciferase | Size (kDa) | Substrate | Requires | Secreted |
| Photinus pyralis | North American firefly luciferase | 61 | D-luciferin | Mg, ATP | No |
| Luciola cruciate | Japanese firefly (Genji-botaru ) luciferase | 64 | D-luciferin | Mg, ATP | No |
| Luciola italic | Italian firefly luciferase | 64 | D-luciferin | Mg, ATP | No |
| Luciola lateralis | Japanese firefly (Heike) luciferase | 64 | D-luciferin | Mg, ATP | No |
| Luciola mingrelica | East European firefly luciferase | 64 | D-luciferin | Mg, ATP | No |
| Photuris pennsylvanica | Pennsylvania firefly luciferase | 64 | D-luciferin | Mg, ATP | No |
| Pyrophorus plagiophthalamus | Click beetle luciferase | 64 | D-luciferin | Mg, ATP | No |
| Phrixothrix hirtus | Railroad worm luciferase | 64 | D-luciferin | Mg, ATP | No |
| Renilla reniformis | Renilla luciferase | 36 | Coelenterazine | N/A | No |
| Rluc8 (mutant of Renillaluciferase) | 36 | Coelenterazine | N/A | No | |
| Green Renilla luciferase | 36 | Coelenterazine | N/A | No | |
| Gaussia princeps | Gaussia luciferase | 20 | Coelenterazine | N/A | Yes |
| Gaussia-Dura luciferase | 20 | Coelenterazine | N/A | Yes | |
| Cypridina noctiluca | Cypridina luciferase | 62 | Vargulin/Cypridina luciferin | N/A | Yes |
| Cypridina hilgendorfii | Cypridina (Vargula) luciferase | 62 | Vargulin/Cypridina luciferin | N/A | Yes |
| Metridia longa | Metridia luciferase | 23.8 | Coelenterazine | N/A | Yes |
| Oplophorus gracilorostris | OLuc | 19 | Coelenterazine | N/A | Yes |
Species-specific luciferase specificity, cofactor requirements and physical characteristics.
Luciferase has gained tremendous popularity in the study of biological systems, because bioluminescence is a sensitive assay and the reagents are easy to use. Examples of areas where bioluminescence is routinely used include:
- In vivo imaging
- Cell proliferation assays
- Protein folding/secretion analyses
- Reporter gene assays
In vivo Imaging
In vivo imaging is a relatively new, non-invasive method that is gaining fast acceptance in biomedical research because of the ability to track molecular events in live animals such as mice and rats. In animal disease models, cells, pathogens, proteins or other molecules are labeled with bioluminescent luciferases and visualized after the localized or systemic addition of substrate. Low-level light is able to pass through tissue, and the level of penetration is dependent upon the wavelength of the emitted light. Therefore, although bioluminescence emitted from deep within an experimental animal can be detected with a sensitive detection system, luciferases that emit light at wavelengths >600 nm have the greatest sensitivity for in vivo imaging applications (2).
Cell viability assays
Bioluminescent assays have also been developed to monitor cell viability and proliferation. Bioluminescent cell viability assays quantitate the free ATP present in metabolically active cells using ATP-dependent luciferases. As a cell population proliferates, the amount of available ATP increases, resulting in a concomitant increase in the bioluminescent signal of the population. Commonly used non-bioluminescent methods to measure cell proliferation include:
- Quantitation of DNA synthesis by measuring titrated thymidine (3H-thymidine) or bromodeoxyuridine (BrdU) uptake
- Identification of dead cells using propidium iodide, a membrane-impermeable DNA-intercalating dye
- Quantitation of the reduction of the intracellular environment by tetrazolium salt reduction (MTT)
- AlamarBlue reduction and quantitation of intracellular ATP concentration.
Reporter gene assays
Reporter gene assays are used to investigate the genetic regulatory elements that control the expression of genes of interest. A gene consists of multiple functional parts, including the coding region that specifies the protein to be made and regulatory elements that control the transcription of the coding region.
The reporter gene assay centers around fusing the putative regulatory elements to a reporter gene and monitoring the amount of the reporter protein expressed. Because reporter expression is under the control of the fused genetic elements, reporter expression is directly correlated with the activity of the regulatory elements. These elements could be promoters, enhancers or 5' or 3' untranslated regions (UTRs) that control either the transcription or translation events in the cell.
The characteristics of a good reporter protein are that it should be easy to assay or detect and is not present normally in the test system. Selection of the reporter system also depends on the experimental system, sensitivity needs and the available detection strategy, be it absorbance, fluorescence or bioluminescence. The proteins that have been traditionally used as reporters include β-galactosidase (lacZ), chloramphenyl acetyltransferase (CAT), β-glucuronidase (GUS) and fluorescent proteins (green, yellow or red fluorescent protein [GFP, YFP or RFP, respectively) and secretory alkaline phosphatase (SEAP).
Luciferase reporter assays
Luciferase assays
Luciferase-based reporter gene assays are widely used because of their ultrasensitive detection capacity and wide dynamic range. These assays involve placing a genetic regulatory element upstream of a luciferase gene and then transferring the resulting reporter construct into animal cells, plant cells or bacteria through transfection, transformation or injection. Expression of the luciferase reporter gene is then measured to quantify the activity of the regulatory element (cis-acting) or proteins (trans-acting) in the biological pathway affected by the target element.
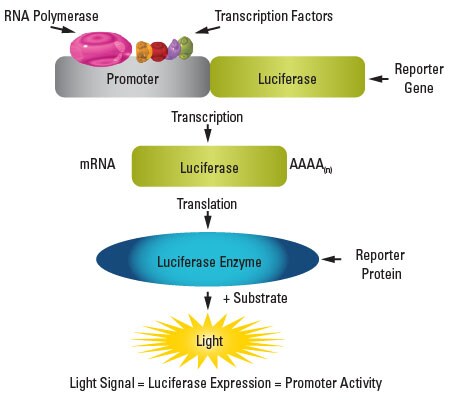
Schematic of the luciferase reporter assay.
Single- vs. dual-spectral assays
Luciferase enzymes can be used as single reporters to study one biological event in a given experiment, but because of their different spectral properties and/or substrates, multiple luciferase enzymes can be combined for multiplex luciferase experiments. With these types of experiments, called dual-spectral luciferase reporter assays, two luciferase enzymes are simultaneously expressed and detected in a single sample. Dual-spectral luciferase reporter assays are ideal for:
- Analyzing more than one target per screen
- Minimizing off-target effects
- Identifying crosstalk between two or more pathways
- Normalizing data to eliminate experimental artifacts
For data normalization experiments, one reporter generally functions as an experimental reporter and the second as an internal control to account for non-specific experimental variations due to operator error or as a result of non-specific effects of biological manipulations or cell treatments. Additional experimental variations include:
- Differences in cell plating
- Differences in cell viability or proliferation due to an experimental treatment
- Edge effects caused by uneven environmental conditions in the cell culture incubator
- Non-specific effects of the treatment compound on the function of the reporter itself
Measurement strategies
Dual-spectral luciferase reporter assays using luciferase enzymes with non-overlapping luminescence spectra can be measured simultaneously by filter-based detection in a single sample using a single reagent. Dual reporter assays like this that use either the same or different substrates are commercially available.
When using luciferases with overlapping spectra, the luciferases can be measured sequentially in two steps using two reagents if they have different substrate specificities. In the two-step method, the luminescence from the first luciferase is measured, and the enzyme is then inactivated. A second reagent containing the substrate for the second luciferase is then added and the activity measured. Theoretically, more than two enzymes can be multiplexed in this way as long as they have different substrate specificities.
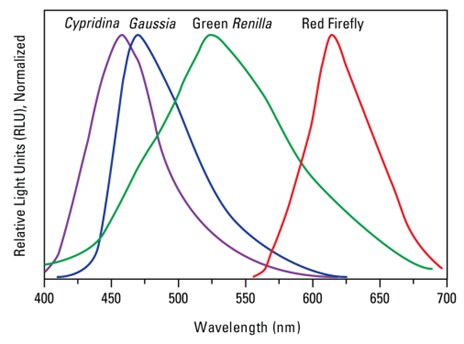
Spectral Resolution between different luciferases. Note: Emission of Red Firefly shifted to the right allowing for multiplexing with other luciferases.
| Luciferase combinations | Multiplex detection strategies | |
| Substrate-based | Spectral/Spatial-based | |
| Native Firefly/Renilla | Luciferin/Coelenterazine | N/A |
| Renilla/Red Firefly | Coelenterazine/Luciferin | 460 nm BP/615 nm LP |
| Green Renilla/Red Firefly | Coelenterazine/Luciferin | 525 nm BP/640 nm LP |
| Gaussia/Renilla | Difficult but possible with coelenterazine analogs | N/A |
| Gaussia/Green Renilla | Difficult but possible with coelenterazine analogs | N/A |
| Gaussia/Firefly | Coelenterazine/Luciferin | N/A |
| Gaussia/Red Firefly | Coelenterazine/Luciferin | 470 nm BP/640 nm LP |
| Gaussia/Cypridina | Coelenterazine/Vargulin | N/A |
| Cypridina/Renilla | Vargulin/Coelenterazine | N/A |
| Cypridina/Green Renilla | Vargulin/Coelenterazine | N/A |
| Cypridina/Red Firefly | Vargulin/Luciferin | 470 nm BP/640 nm LP |
| Cypridina/Gaussia/any Firefly | Vargulin/Coelenterazine/Luciferin | N/A |
| Cypridina/any Renilla/Firefly | Vargulin/Coelenterazine/Luciferin | N/A |
| BP: band pass; LP: long pass | ||
Dual-spectral luciferase assay combinations and multiplex detection strategies.
Time-course assays using dual-spectral luciferase assays
Dual-spectral luciferase assays are ideal for performing time-course assays using a secreted luciferase to measure the experimental promoter activity and an intracellular luciferase under the control of a housekeeping gene promoter to measure cell viability. Experimental promoter activity results in luciferase secretion into the culture medium, which is collected at various time points throughout the experiment. The cells are then lysed at the completion of the time-course, and the lysate is measured for intracellular luciferase activity as the control for cell viability.
Not all commercially available dual-spectral luciferase assays are compatible with time-course assays because of the requirement for a secreted luciferase reporter. For example, Renilla luciferase is not secreted, and thus the Pierce Renilla-Firefly Luciferase Dual Reporter Assay Kit will not work for this type of time-course assay. However, Cypridina and Gaussia luciferases are secreted; therefore, Pierce Cypridina-Firefly and Gaussia-Firefly dual-spectral luciferase kits are compatible with time-course assays.
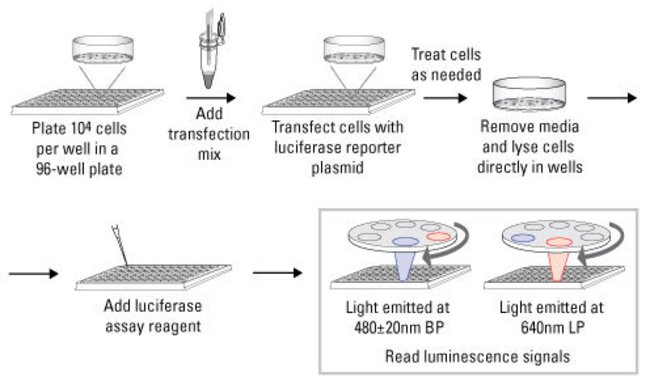
Workflow for the Cypridina-Firefly Luciferase Dual Assay Kit
Recommended reading
- Keilin D. (1966) The history of cell respiration and cytochrome. Cambridge, Cambridge U.P. xx, 416.
- Negrin R. S. and Contag C. H. (2006) In vivo imaging using bioluminescence: A tool for probing graft-versus-host disease. Nat Rev Immunol. 6, 484-90.
- Smith K. C. (1989) the science of photobiology. New York: Plenum Press. Viii, 426.
- Campbell A. K. and Herring P. J. (1990) Imidazolopyrazine bioluminescence in copepods and other marine organisms. Marine Biology. 104, 219-25.
- Robison B. H. and Young R. E. (1981) Bioluminescence in pelagic octopods. Pacific Science. 35, 39-44
For Research Use Only. Not for use in diagnostic procedures.