Search
siRNA Libraries Tie Genes to Cell Functions
Ambion has developed human siRNA libraries that target important gene classes including kinases, ion channels, receptor classes, molecular motors, and many others (Figure 1). Applying these siRNA libraries to cell culture screens reveals human genes that are critical for cellular functions. The diversity of cell functions that can be characterized using siRNA libraries is limited only by the range of phenotypic assays that can be developed. With siRNA libraries, you will be able to identify critical human genes faster and easier than ever before.
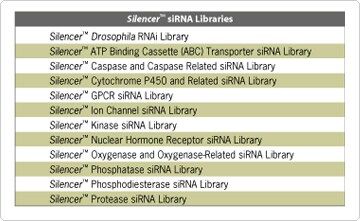
Figure 1. Ambion siRNA Libraries.
Scientists at Ambion have already used several Silencer™ siRNA libraries in quantitative assays to identify genes involved in apoptosis, cell proliferation, TRAIL induction, and p53 activation. Below we provide a general overview of an effective screening process.
Overview of the Screening Process
(1) Develop a quantitative assay to monitor cellular process being studied.
Assays that measure the intensity of a cellular phenotype range from microscopic assays that monitor cell size, cell cycle status, or antibody staining; to enzymatic assays that assess the turnover of a specific substrate in a cell lysate; to direct measurements of biomolecules or small molecules in lysates, on cells, or in medium.
A successful screen must use an assay that quantitates a cellular phenotype and maximizes the signal-to-noise ratio. An effective assay must include quality positive and negative control siRNAs. The positive control siRNA should produce the desired phenotype when transfected into cells, and the target of the positive control siRNA should ideally be a gene that is known to act in the process being studied. For instance, if the goal is to identify genes that are involved in a signaling pathway after stimulation by a specific molecule then transfecting an siRNA targeting the receptor for the moleule should disrupt the cell signaling and result in the desired phenotype. Likewise, a negative control siRNA should have essentially no effect on the assay results, which should be comparable to untransfected or mock-transfected cells.
Maximizing signal-to-noise involves testing variables like assay time, assay components (i.e. the reporter), cell type, and length of time between transfection and assay. The greater the difference in assay results between the positive and negative controls, the greater the spread will be in the screening results, and thus, the better the opportunity to identify interesting genes.
(2) Optimize transfection conditions for the desired cells.
The first step is to identify a transfection reagent and plating conditions that maximize uptake of active siRNA while maintaining high cell viability. We find it useful to measure both knockdown and cell viability in cells transfected with 2-5 different transfection reagents when using cell lines, or 5-10 electroporation conditions (e.g. varying cell concentration, voltage, pulse length, pulse number, siRNA concentration etc.) when using primary or suspension cells. Transfection can then be optimized for the reagent or electroporation condition that worked best among the conditions tested.
Ambion has several transfection agents developed for effective siRNA transfer to specific cell types; see siPORT™ siRNA Electroporation Buffer for more information.
Screening siRNAs targeting hundreds of genes requires conditions for high-throughput transfection. siPORT™ NeoFX™ facilitates the transfection of up to 1,000 wells in less than an hour without the need for robotics.
(3) Screen
Once the assay and transfection processes have been optimized, a library of siRNAs can be introduced sequentially into cells in a 24 or 96 well plate. Triplicate transfections for each siRNA provide enough data for reasonable statistical analysis. Positive and negative control siRNAs transfected on each plate provide quantitative numbers that will allow data from different plates to be normalized and will provide the range of phenotypes that can be used to identify genes that rate as "hits".
Both positive and negative control siRNAs are available from Ambion.
(4) Validate hits
"Hits" are genes for which an siRNA produces a phenotype that is similar to the positive control. Validating a hit involves showing that the observed phenotype is due to reducing the expression of the target gene. We confirm a hit by individually transfecting three distinct siRNAs targeting the same gene, monitoring reduction in the target mRNA or protein, and then measuring the phenotype. In theory, all siRNAs that similarly reduce the expression of a target gene should yield the same phenotype. In practice, we find occasional off-target sequence effects of siRNAs that lead to false positive or false negative phenotypes for a given siRNA. The use of three individual siRNAs essentially eliminates false positives and false negatives.
Assays that measure the intensity of a cellular phenotype range from microscopic assays that monitor cell size, cell cycle status, or antibody staining; to enzymatic assays that assess the turnover of a specific substrate in a cell lysate; to direct measurements of biomolecules or small molecules in lysates, on cells, or in medium.
A successful screen must use an assay that quantitates a cellular phenotype and maximizes the signal-to-noise ratio. An effective assay must include quality positive and negative control siRNAs. The positive control siRNA should produce the desired phenotype when transfected into cells, and the target of the positive control siRNA should ideally be a gene that is known to act in the process being studied. For instance, if the goal is to identify genes that are involved in a signaling pathway after stimulation by a specific molecule then transfecting an siRNA targeting the receptor for the moleule should disrupt the cell signaling and result in the desired phenotype. Likewise, a negative control siRNA should have essentially no effect on the assay results, which should be comparable to untransfected or mock-transfected cells.
Maximizing signal-to-noise involves testing variables like assay time, assay components (i.e. the reporter), cell type, and length of time between transfection and assay. The greater the difference in assay results between the positive and negative controls, the greater the spread will be in the screening results, and thus, the better the opportunity to identify interesting genes.
(2) Optimize transfection conditions for the desired cells.
The first step is to identify a transfection reagent and plating conditions that maximize uptake of active siRNA while maintaining high cell viability. We find it useful to measure both knockdown and cell viability in cells transfected with 2-5 different transfection reagents when using cell lines, or 5-10 electroporation conditions (e.g. varying cell concentration, voltage, pulse length, pulse number, siRNA concentration etc.) when using primary or suspension cells. Transfection can then be optimized for the reagent or electroporation condition that worked best among the conditions tested.
Ambion has several transfection agents developed for effective siRNA transfer to specific cell types; see siPORT™ siRNA Electroporation Buffer for more information.
Screening siRNAs targeting hundreds of genes requires conditions for high-throughput transfection. siPORT™ NeoFX™ facilitates the transfection of up to 1,000 wells in less than an hour without the need for robotics.
(3) Screen
Once the assay and transfection processes have been optimized, a library of siRNAs can be introduced sequentially into cells in a 24 or 96 well plate. Triplicate transfections for each siRNA provide enough data for reasonable statistical analysis. Positive and negative control siRNAs transfected on each plate provide quantitative numbers that will allow data from different plates to be normalized and will provide the range of phenotypes that can be used to identify genes that rate as "hits".
Both positive and negative control siRNAs are available from Ambion.
(4) Validate hits
"Hits" are genes for which an siRNA produces a phenotype that is similar to the positive control. Validating a hit involves showing that the observed phenotype is due to reducing the expression of the target gene. We confirm a hit by individually transfecting three distinct siRNAs targeting the same gene, monitoring reduction in the target mRNA or protein, and then measuring the phenotype. In theory, all siRNAs that similarly reduce the expression of a target gene should yield the same phenotype. In practice, we find occasional off-target sequence effects of siRNAs that lead to false positive or false negative phenotypes for a given siRNA. The use of three individual siRNAs essentially eliminates false positives and false negatives.