Search
To enable autophagy studies, we offer specific tools to report LC3B localization: an antibody against LC3B, and Premo™ sensors. The Premo™ Autophagy Sensors are genetically encoded constructs expressed as either GFP or RFP fused to the N-terminus of LC3B, with the C-terminus free for cleavage and subsequent recruitment of autophagosomes.
Imaging Autophagy in Physiologically Relevant Models
Aberrant autophagy is implicated in many important disease states such as cancer, neurodegeneration, and immunological diseases [2], and therefore small molecules that can remedy these deficiencies hold great therapeutic promise [3]. With this in mind, it is important to study autophagy in physiologically relevant cell models such as primary cells and neurons. The Premo™ Autophagy Sensor LC3B leverages BacMam 2.0 delivery technology to yield high-efficiency transduction of primary cells, stem cells, and neurons in a simple, one-step protocol (Figure 1; also see the article on Transforming Live-Cell Microscopy with CellLight® Reagents). The ability to study autophagosome formation and other aspects of autophagy, and to observe the effects of small molecules and gene manipulation on them in physiologically relevant cell models, is of great importance, not only in understanding autophagy but also in producing effective treatments.
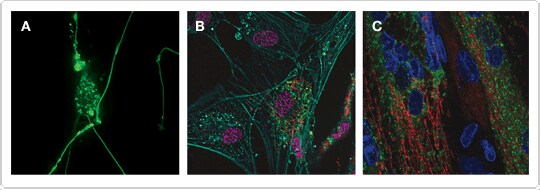
Figure 1. Expression of the Premo™ Autophagy Sensor LC3B-GFP in primary cell types. Rat hippocampal neurons (A), human dermal fibroblasts (B), and human skeletal muscle cells (C) were transduced with the Premo™ Autophagy Sensor LC3B-GFP. This autophagy sensor can be easily combined with other fluorophores to obtain a multiparametric readout of autophagy. In B, the fibroblasts were also transduced with CellLight® Mitochondria-RFP and stained with Hoechst 33342 and Alexa Fluor® 647 phalloidin to label mitochondria (red), nuclei (magenta), and filamentous actin (cyan), respectively. In C, the cells were transduced with CellLight® Mitochondria-RFP and stained with Hoechst 33342 (blue).
Autophagic Flux
Western blot analysis using anti-GFP can be used to detect autophagic flux. Under resting conditions, anti-GFP detects a 45 kDa band (representing the 27 kDa GFP and 18 kDa LC3B). If autophagy is induced, the Premo™ Autophagy Sensor LC3B is recruited to the autophagosome, which then fuses with the lysosome to form the autolysosome. Once in the autolysosome, the LC3B portion of the sensor is more prone to degradation than GFP, resulting in the detection of only a 27 kDa band representing free GFP [4].
Involvement of Other Cellular Organelles
Invitrogen offers organic dyes and fluorescent proteins targeted to mitochondria, the ER, peroxisomes, and lysosomes that complement studies of autophagy. The combination of these probes with autophagy-specific markers such as the Premo™ Autophagy Sensor LC3B-GFP or Premo™ Autophagy Sensor LC3B-RFP allows multicolor imaging of the interaction of other cellular components with autophagosomes during their formation and fusion with lysosomes to form autolysosomes [5,6] (Figure 2). This last step is critical in assessing the degree to which autophagy will be completed. In addition to lysosome-targeted fluorescent proteins and LysoTracker® dyes, fluorescent dextran molecules can be used to label lysosomes [7].
To visualize the formation of autolysosomes, it may be beneficial to use the Premo™ Autophagy Sensor LC3B-RFP rather than LC3B-GFP, as the pKa of TagRFP is lower than that of GFP. This lower pKa renders the TagRFP-based sensor more suitable for visualizing autolysosome formation, as its fluorescence is less likely than that of emGFP to be quenched by the acidic environment of the autolysosomes [8].
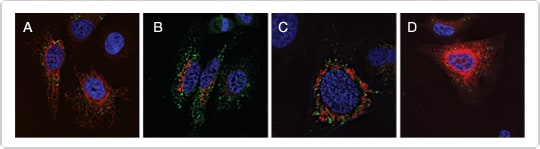
Figure 2. Multicolor imaging of autophagosomes and other organelles. A549 cells were transduced with the Premo™ Autophagy Sensor LC3B-GFP and CellLight® reagents to label specific organelles. At 24 hr after transduction, cells were treated with 50 µM chloroquine overnight to favor autophagosome formation. Nuclei were labeled with Hoechst 33342 (blue). Cells were transduced with (A) CellLight® Mitochondria-RFP, (B) CellLight® Golgi-RFP, (C) CellLight® Lysosomes-GFP, and (D) CellLight® ER-RFP.
Tracking Autophagy From Start to Finish
The tools and techniques outlined above allow the highly dynamic processes occurring during autophagy, from autophagosome formation to degradation in the autolysosome, to be studied in a relevant cellular setting. Learn more about products for research on autophagy.
- Klionsky DJ (2007) Nat Rev Mol Cell Biol 8:931–937.
- Mizushima N, Levine B, Cuervo AM et al. (2008) Nature 451:1069–1075.
- Rubinsztein DC, Gestwicki JE, Murphy LO et al. (2007) Nat Rev Drug Discov 6:304–312.
- Mizushima N, Yoshimori T, Levine B (2010) Cell 140:313–326.
- Hailey DW, Rambold AS, Satpute-Krishnan P et al. (2010) Cell 141:656–667.
- Yu L, McPhee CK, Zheng L et al. (2010) Nature 465:942–946.
- Saftig P, Klumperman J (2009) Nat Rev Mol Cell Biol 9:623–635.
- Kimura S, Noda T, Yoshimori T (2007) Autophagy 3:452–460.
For Research Use Only. Not for use in diagnostic procedures.
See a complete listing of the products discussed in this article.
View products
Find more information on LC3B protein detection.
- Learn More about Tools to Investigate Autophagy
Get a copy of this article as it appears in the print version of BioProbes 63.
