Search
Automating RNA Isolation
Large scale gene expression screening projects require reliable high-throughput methods to isolate high quality RNA from a large number of samples. To meet this growing need, Ambion developed the
RNAqueous™-96 Kit. This kit, which utilizes an RNA-binding glass-fiber filter, provides high yields of intact RNA in a 96-well plate format. An optional on-the-filter DNase treatment can be performed to ensure removal of genomic DNA for RT-PCR applications. Because the kit is compatible with commonly used 96-well plate vacuum manifolds, the procedure is readily automated for use with high throughput liquid handling systems. Recently, Ambion, in collaboration with PerkinElmer Life Sciences, developed an automation protocol for the RNAqueous-96 Kit using the Packard MultiPROBE® II HT Liquid Handling System. This method reproducibly yields high quality total RNA from both adherent and suspended mammalian cultured cells.
Reproducible Isolation of High Quality RNA
To demonstrate the high quality of RNA that can be obtained using the RNAqueous-96 Kit on a robotic platform, RNA from multiple HeLa S3 and K562 cell samples was isolated using the method described in "RNAqueous-96 Automation Protocol". Briefly, HeLa S3 (adherent) and K562 (suspended) cells were transferred to a deep well 96-well plate at 3.75 x 105 cells/well. HeLa cells were incubated for 2 days prior to RNA isolation, whereas K562 cells were processed immediately. The resulting RNA was assessed for both yield and quality on an Agilent 2100 bioanalyzer. Yields were consistently high (~5 pg per cell for HeLa S3 cells and ~8 pg per cell for K562 cells) and RNA samples were uniformly intact. Furthermore, genomic DNA was effectively removed. An electropherogram demonstrating the high quality of the RNA recovered is shown in Figure 1.
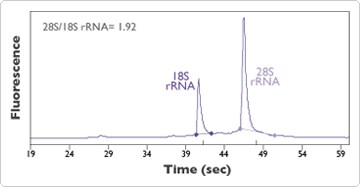
Figure 1. RNA Isolated Using the Automated RNAqueous-96 Protocol. Cultured K562 cells (human leukemia, grown in suspension) were pelleted and resuspended in 1X phosphate buffered saline (PBS), counted and transferred to 48 wells of a deep well 96-well plate at 3.75 x 105 cells/well. Total RNA was isolated using the automated method described at "RNAqueous-96 Automation Protocol", using RNAqueous-96 and a MultiPROBE II HT Liquid Handling System with a Gripper Integration Platform. This RNA sample, which was representative of the 48 samples isolated and analyzed, has a 28S rRNA to 18S rRNA ratio of 1.92, indicating a high level of integrity. This isolation yielded 24 µg of RNA (344 ng/µl in 70 µl).
Lack of reproducibility can be an issue with some RNA isolation methods. Real-time RT-PCR was used to assess the reproducibility of the automated RNAqueous-96 procedure. Because real-time RT-PCR is quantitative, reproducible, and can be used to analyze multiple samples simultaneously, it is a good choice for such an analysis. Both an abundant (GAPDH) and a rare target (hTERT: catalytic subunit of human telomerase) were individually analyzed in the previously described HeLa S3 and K562 RNA samples using a TaqMan® assay. Results from the analysis of 9 samples from K562 cells are shown in Figure 2. Cycle threshold (Ct) values, which reflect the amount of target RNA present in the sample, were calculated (Figure 3). Although some variation in yield between replicate wells was observed, it is most likely due to several factors unrelated to the isolation method, including variation in the number of cells manually dispensed to each well, and variation in cell growth and turnover during incubation. Importantly, the consistent ratio of hTERT to GAPDH in replicate samples indicates that both high abundance and low abundance mRNAs are reproducibly isolated with the automated RNAqueous-96 protocol.
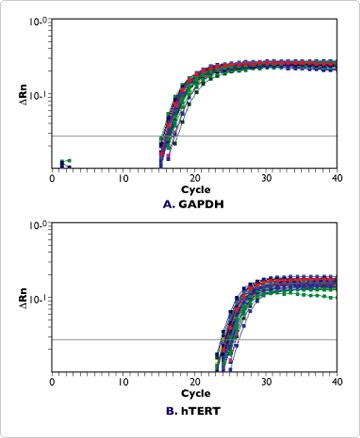
Figure 2. Real-time RT-PCR Detection of GAPDH and hTERT in Replicate Samples Isolated Using the RNAqueous-96 Automation Protocol. RNA was isolated as described in Figure 1. Samples were analyzed by real-time RT-PCR using primers and a TaqMan® probe for GAPDH (A.) or hTERT (B.) in an ABI 7700 Sequence Detector. Data from 9 samples, each indicated by a different color line, are shown here. The average cycle threshold (Ct) values for all of the samples are shown in Figure 3.
Figure 3. Average Ct Values from Real-time RT-PCR Analysis of 48 Samples.
Although this example used the Packard MultiPROBE II HT Liquid Handling System, the RNAqueous-96 Kit is readily adaptable to other automated platforms. The RNAqueous-96 Kit includes reagents and supplies to perform 192 isolations, from 100 to 2 x 106 cells or 0.1 to 1.5 mg of tissue per sample. Reagents for an optional on-the-filter DNase treatment are included.
MultiPROBE® is a registered trademark of Packard BioScience Company. TaqMan® is a registered trademark of Roche Molecular Systems, Inc.

Figure 1. RNA Isolated Using the Automated RNAqueous-96 Protocol. Cultured K562 cells (human leukemia, grown in suspension) were pelleted and resuspended in 1X phosphate buffered saline (PBS), counted and transferred to 48 wells of a deep well 96-well plate at 3.75 x 105 cells/well. Total RNA was isolated using the automated method described at "RNAqueous-96 Automation Protocol", using RNAqueous-96 and a MultiPROBE II HT Liquid Handling System with a Gripper Integration Platform. This RNA sample, which was representative of the 48 samples isolated and analyzed, has a 28S rRNA to 18S rRNA ratio of 1.92, indicating a high level of integrity. This isolation yielded 24 µg of RNA (344 ng/µl in 70 µl).
Lack of reproducibility can be an issue with some RNA isolation methods. Real-time RT-PCR was used to assess the reproducibility of the automated RNAqueous-96 procedure. Because real-time RT-PCR is quantitative, reproducible, and can be used to analyze multiple samples simultaneously, it is a good choice for such an analysis. Both an abundant (GAPDH) and a rare target (hTERT: catalytic subunit of human telomerase) were individually analyzed in the previously described HeLa S3 and K562 RNA samples using a TaqMan® assay. Results from the analysis of 9 samples from K562 cells are shown in Figure 2. Cycle threshold (Ct) values, which reflect the amount of target RNA present in the sample, were calculated (Figure 3). Although some variation in yield between replicate wells was observed, it is most likely due to several factors unrelated to the isolation method, including variation in the number of cells manually dispensed to each well, and variation in cell growth and turnover during incubation. Importantly, the consistent ratio of hTERT to GAPDH in replicate samples indicates that both high abundance and low abundance mRNAs are reproducibly isolated with the automated RNAqueous-96 protocol.

Figure 2. Real-time RT-PCR Detection of GAPDH and hTERT in Replicate Samples Isolated Using the RNAqueous-96 Automation Protocol. RNA was isolated as described in Figure 1. Samples were analyzed by real-time RT-PCR using primers and a TaqMan® probe for GAPDH (A.) or hTERT (B.) in an ABI 7700 Sequence Detector. Data from 9 samples, each indicated by a different color line, are shown here. The average cycle threshold (Ct) values for all of the samples are shown in Figure 3.
| Cell Line | Target | Average Ct Value | Standard Deviation |
|---|---|---|---|
| K562 | GAPDH | 15.86 | 0.45 |
| hTERT | 23.63 | 0.54 | |
| hTERT/GAPDH | 1.49 | 0.03 (CV=1.79%) | |
| HeLA S3 | GAPDH | 16.18 | 0.8 |
| hTERT | 26 | 1.01 | |
| hTERT/GAPDH | 1.61 | 0.30 (CV=2.1%) |
Figure 3. Average Ct Values from Real-time RT-PCR Analysis of 48 Samples.
Although this example used the Packard MultiPROBE II HT Liquid Handling System, the RNAqueous-96 Kit is readily adaptable to other automated platforms. The RNAqueous-96 Kit includes reagents and supplies to perform 192 isolations, from 100 to 2 x 106 cells or 0.1 to 1.5 mg of tissue per sample. Reagents for an optional on-the-filter DNase treatment are included.
MultiPROBE® is a registered trademark of Packard BioScience Company. TaqMan® is a registered trademark of Roche Molecular Systems, Inc.