Search
RNA quantitation is an important and necessary step prior to most RNA analysis methods
Here we discuss three common methods used to quantitate RNA and tips for optimizing each of these methods.
UV spectroscopy
The traditional method for assessing RNA concentration and purity is UV spectroscopy. The absorbance of a diluted RNA sample is measured at 260 and 280 nm. The nucleic acid concentration is calculated using the Beer-Lambert law, which predicts a linear change in absorbance with concentration (Figure 1).
Using this equation, an A260 reading of 1.0 is equivalent to ~40 µg/ml single-stranded RNA. The A260/A280 ratio is used to assess RNA purity. An A260/A280 ratio of 1.8 2.1 is indicative of highly purified RNA.

Figure 1. Beer-Lambert Law for calculating UV absorbance by nucleic acid.
UV spectroscopy is the most widely used method to quantitate RNA. It is simple to perform, and UV spectrophotometers are available in most laboratories. The method does have several drawbacks, but they can be minimized by following these tips:
Tips for optimizing performance
- Because this method does not discriminate between RNA and DNA, it is advisable to first treat RNA samples with RNase-free DNase to remove contaminating DNA.
- Other contaminants such as residual proteins and phenol can interfere with absorbance readings, so care must be taken during RNA purification to remove them.
- Sample readings are made in quartz cuvettes. Dirty cuvettes and dust particles cause light scatter at 320 nm which can impact absorbance at 260 nm. Since neither proteins nor nucleic acids absorb at 320 nm, perform a background correction by making readings from a blank (diluent only) at 320 nm, as well as 260 nm and 280 nm.
- The A260/A280 ratio is dependent on both pH and ionic strength. As pH increases, the A280 decreases while the A260 is unaffected. This results in an increasing A260/A280 ratio (Wilfinger, et. al 1997). Because water often has an acidic pH, it can lower the A260/A280 ratio. We recommend using a buffered solution with a slightly alkaline pH, such as TE (pH 8.0), as a diluent (and as a blank) to assure accurate and reproducible readings. An example of the variation in A260/A280 ratio at different pH values is shown in Table 1.
- Make sure your RNA dilution is within the linear range of your spectrophotometer. Usually absorbance values should fall between 0.1 and 1.0. Solutions that are outside this range cannot be measured accurately. Generally the greatest error occurs at lower concentrations.
- Because an A260 of 0.1 corresponds to ~4 µg/ml RNA, it is often impractical to use UV spectroscopy to quantitate RNA isolated from small samples that will have lower concentrations once diluted. Fortunately, there are alternative methods for accurately quantitating small amounts of RNA two are described below.
Table 1. Effects of pH on A260/A280 ratio.
| Blank/Diluent | A260/A280 ratio |
|---|---|
| DEPC-treated water (pH 5-6) | 1.60 |
| Nuclease-free water (pH 6-7) | 1.85 |
| TE (pH 8.0) | 2.14 |
The Agilent 2100 bioanalyzer uses a combination of microfluidics, capillary electrophoresis, and fluorescent dye that binds to nucleic acid to evaluate both RNA concentration and integrity. An RNA reference standard (the RNA 6000 Ladder Cat# 7152; Ambion) and a microfluidics chip (The RNA Lab Chip; Agilent Technologies) are also required. The RNA 6000 Ladder is composed of six RNAs ranging in size from 0.2 6 kb. The ladder and samples are loaded in designated wells on the RNA Lab Chip. Size and mass information is provided by the fluorescence of RNA molecules as they move through the channels of the chip. The instrument software automatically compares the peak areas from unknown RNA samples to the combined area of the six RNA 6000 Ladder RNA peaks to determine the concentration of the unknown samples. The RNA 6000 Nano System has a broad dynamic range and can quantitate between 25 500 ng/ml of RNA with a covariance of ~10%.
Perhaps the most powerful feature of the Agilent 2100 bioanalyzer is its ability to provide information about RNA integrity. As each RNA sample is analyzed, the software generates both a gel-like image and an electropherogram (Figure 2). When analyzing total RNA, the areas under the 18S and 28S ribosomal RNA peaks are used to calculate the ratio of these two major ribosomal RNA species and these data are displayed along with quantitation data on individual electropherograms (Figure 2a). Significant changes in the ratios of the 18S and 28S ribosomal RNA peaks are indicative of degraded RNA.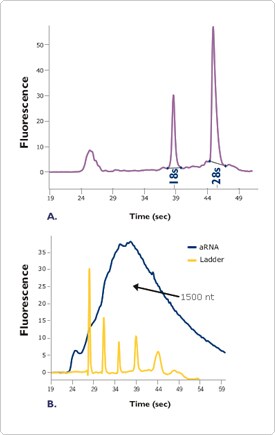 Figure 2. Agilent 2100 Bioanalyzer Electropherograms of RNA Samples.
Figure 2. Agilent 2100 Bioanalyzer Electropherograms of RNA Samples.
A. Electropherogram of a Total RNA Sample. Total RNA (100 ng) was analyzed on an Agilent 2100 bioanalyzer. The resulting electropherogram shows the characteristic signature of a high quality total RNA sample.
B. Electropherogram of Amplified aRNA Sample. Total RNA 2 µg corresponding to 60 ng mRNA) was amplified using the MessageAmp aRNA Kit (Ambion Cat# 1750) resulting in (90 µg aRNA, a 1500 fold amplification. The aRNA (900 ng) was analyzed on an Agilent 2100 bioanalyzer. The resulting electropherogram shows the classic output of a high quality aRNA sample.
In addition to its usefulness for analysis of total RNA, the bioanalyzer is also a superior tool for analyzing mRNA and amplified aRNA (antisense RNA) integrity. Intact mRNA and aRNA profiles consist of a broad distribution of signal, with the bulk of the RNA usually falling between 1 and 2 kb, though this will vary from tissue to tissue (Figure 2b). A significant shift of the profile towards lower molecular weights is indicative of poor RNA integrity.
Tips for Optimizing Performance
- The area of the peaks derived from the RNA 6000 Ladder is used as a mass standard for unknowns, so accurate quantitation of your unknowns is dependent on careful handling of this standard. We recommend that the RNA 6000 Ladder be thoroughly mixed and carefully pipetted to reduce error. For best performance, the standard should be aliquoted into non-stick, nuclease-free tubes to avoid multiple freeze-thaw cycles of a single stock tube.
- Quantitation is affected by ionic strength of the sample, which can quench fluorescence in RNA samples. Therefore, when possible, RNA should be suspended in nuclease-free water to minimize differences between the RNA 6000 Ladder and the sample to be measured. If this is not possible, be aware that the unknown concentration may be underestimated.
- Generally, we find that some 23S and 28S rRNAs do not migrate according to their molecular weights. For example, mammalian 28S rRNA, 4.8 kb in length, consistently migrates just ahead of the 4 kb peak in the RNA 6000 Ladder. This is likely due to the highly structured nature of 23S and 28S rRNAs.
- Although this assay has a broad linear range (~25 ng 500 ng) overloading the chip with RNA can affect performance of the RNA Lab Chip. For consistent results, we recommend loading 50 ng 250 ng of RNA.
- The fluorescent dye is light sensitive, so store dye concentrate and working solutions away from light; e.g. wrap tubes in foil.
- Follow the manufacturer's recommendations for maintenance of the electrodes and the priming station. Poor Lab Chip Loading (priming) and formation of salt bridges between electrodes are common causes of poor assay performance.
Reference
For Research Use Only. Not for use in diagnostic procedures.