Search
Laser capture microdissection (LCM)
Obtaining specific populations of cells from heterogeneous samples is a major challenge in studying gene expression profiles of specific cell types. Laser capture microdissection (LCM), developed in a collaborative effort between the NIH and Arcturus, Inc. specifically to isolate cancer cells from normal tissue, can be used to separate individual cell types from within complex tissues. RNA can then be extracted from these microdissected cells to analyze their gene expression.
How LCM works
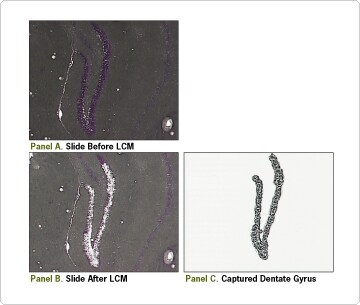
LCM can capture cells or discrete morphological structures from thin tissue sections. Frozen, O.C.T.-embedded tissue sections are prepared for the LCM process by fixation, staining, and dehydration. Sections are visualized through a thermoplastic film attached to the bottom of a microcentrifuge tube cap. A laser pulse is directed through the film onto the target cells. The plastic film melts onto the targeted area, then cools and bonds with the underlying cells. The film, along with the adhered target cells, is collected (Figure 1). RNA can be isolated from these captured cells for real-time PCR and mRNA expression profiling. (See our tips article Getting Intact RNA from LCM Samples.)
Capture and analyses of dentate gyrus from mouse brain
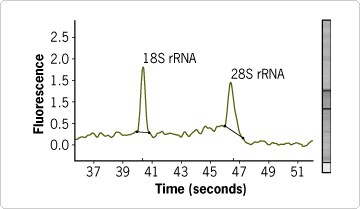
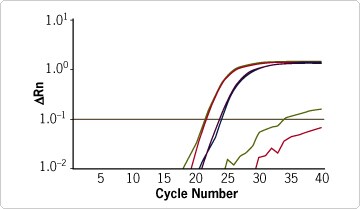
LCM was used to separate dentate gyrus from mouse brain sections (Figure 1). RNA from the captured dentate gyrus was purified using the RNAqueous®-Micro Kit (see sidebar) and the purified RNA was assessed on an Agilent bioanalyzer (Figure 2). This RNA was of sufficient quality and amount for real-time RT-PCR analysis, which showed consistency between replicates and very low DNA contamination in the RNA samples purified with RNAqueous-Micro Kit (Figure 3).
Purification of total RNA from micro-sized samples, such as those obtained by laser capture microdissection, needle biopsies, and fine dissection, as well as samples comprised of small numbers of cultured cells, can be a challenge. In these cases, it is desirable to recover the total RNA in a small volume so that the entire sample can be used in downstream applications, such as reverse transcription. Keeping the sample as concentrated as possible also facilitates analysis and quantitation of the total RNA using sensitive microfluidic assays (e.g. Agilent 2100 bioanalyzer).
Ambion's RNAqueous®-Micro Kit was developed to optimize recovery of highly concentrated total RNA from micro-sized samples. The RNAqueous-Micro Kit protocol begins with sample disruption in a chaotropic solution to lyse cells and inactivate cellular nucleases. The cell lysate is loaded onto a glass fiber filter washed, and the RNA eluted. The filter cartridge provided in the RNAqueous-Micro Kit is only a few millimeters in diameter so that a small volume of fluid is enough to completely saturate it. This allows the RNA to be efficiently eluted in as little as 20 µl. After elution, the RNA is treated with DNase I to eliminate genomic DNA contamination that can interfere with RT-PCR assays. The DNase is then removed using the DNA-free™ Removal Reagent which removes the DNase I and buffer ions in a simple, quick step without organic extraction or heat inactivation. (Heat inactivation of DNase can cause divalent cation-mediated degradation of the RNA.) Using the RNAqueous-Micro Kit, the entire RNA isolation procedure, including DNase treatment, takes about 30 minutes.