Search
Isolating miRNA for Profiling Studies
How to Capture miRNAs
Isolation of miRNA begins when total RNA that includes the small RNA fraction is isolated from the samples of interest. However, not all isolation methods retain the small RNA fraction, resulting in loss of miRNAs. Therefore it is important to use RNA isolation methods specifically adapted to retain it. Ambion provides a complete line of RNA isolation kits that allow purification of the total RNA fraction, including small RNAs, from fresh and frozen tissue, from blood, and from formaldehyde- or paraformaldehyde-fixed, paraffin-embedded (FFPE) tissues. Use any of these sources of miRNA containing total RNA for qRT-PCR or Northern analysis. For miRNA profiling by microarray, we recommend that the mature miRNA fraction be purified from any of these total RNA samples and then labeled for microarray analysis.
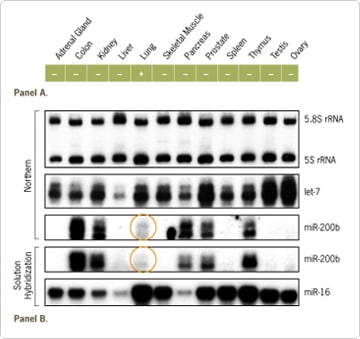
Figure 1. Detection of microRNAs. Not all isolation methods retain the small RNA fraction, resulting in loss of miRNAs. Likewise, not all commercially available total RNAs contain small RNAs. In the report by Grad et al. [Grad et al. (2003) Mol Cell11:1253–1263.], total RNA from a commercially available source was used to detect the presence of miR-200b in various human tissues (by Northern blotting). In this study, miR-200b expression was only detected in human lung tissue (summary of data shown in Panel A). However, it was possible to detect miR-200b in several human tissues using both Northern blot and solution hybridization assay when the total RNA (Ambion’s FirstChoice® Total RNA) was prepared using a method that ensured small RNA retention (B). Interestingly, the levels of miR-200b detected in the lung (the tissue in which miR-200b was previously detected) were lower than that of several other tissues (colon, kidney, pancreas, prostate, and thymus).
Initial Isolation of Total RNA Containing miRNAs: mirVana™ miRNA Isolation Kit and mirVana™ PARIS™ Kit
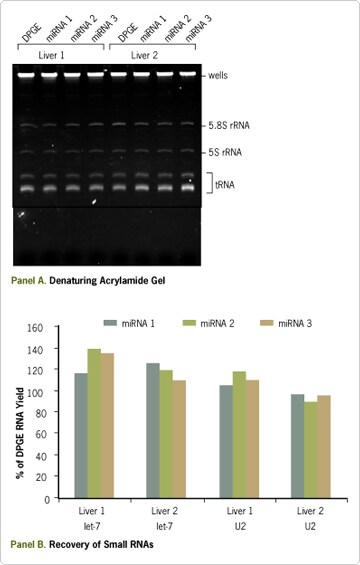
Figure 2. mirVana™ miRNA Isolation Kit for Efficient Recovery of miRNA. (A) Total RNA was isolated from the same mouse liver lysate using a double phenol/guanidinium extraction (DPGE) or the mirVana miRNA Isolation Kit procedure in triplicate (miRNA 1 to 3). The experiment was performed with two different mouse liver lysates. Each sample RNA (1 µg) was analyzed on a denaturing 15% polyacrylamide gel stained with ethidium bromide. (B) RNAs from the same gel were transferred to a membrane and probed for U2 snRNA and let-7 miRNA. The relative amount of small RNA in each lane was quantified with a phosphorimager. The graph shows the percentage of recovery with respect to the DPGE prep.
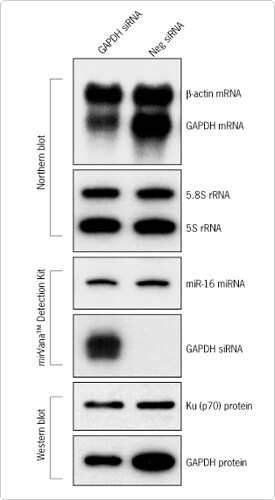
The mirVana PARIS Kit permits quantitative recovery of both native protein and all RNA species, including small RNAs, from the same sample (Figure 3). Samples are first homogenized with a special Cell Disruption Buffer that includes nonionic detergent to extract intact protein; a portion of the lysate can be used directly for common applications such as Western blotting, two-dimensional gel electrophoresis, or enzymatic assays. RNA is isolated from the remainder of the lysate using a procedure similar to the mirVana miRNA Isolation Kit.
The small RNA containing total RNA fraction from either the mirVana miRNA Isolation Kit or the mirVana PARIS Kit (or other sources) can be further enriched using the protocol and reagents included in both kits. The total RNA or enriched miRNA is ideal for qRT-PCR and other miRNA detection techniques.
Ambion also has several isolation products for obtaining total RNA that includes the miRNA fraction from special samples. These include kits for isolating RNA from extremely small samples such as LCM tissue, from the leukocyte fraction of whole blood, and from FFPE treated samples. See sidebar, Isolating miRNAs from Special Samples.
Now that I have my miRNA?
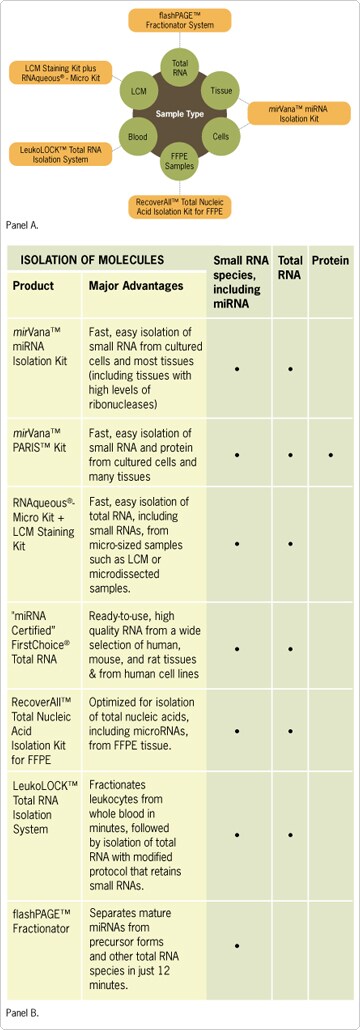
Figure 6. Selection Guide for Small RNA Isolation and Enrichment.
RNAqueous®-Micro + LCM Staining Kit
The RNAqueous-Micro Kit is optimized for the purification of total RNA from micro sized samples such as laser captured microdissected (LCM) samples, microdissected structures, and tissue punches. A simple modification in the protocol allows the user to recover both large and small RNA species including tRNAs, 5S rRNA, and microRNAs. The RNA is eluted in a minimal volume of 20 µl and residual genomic DNA is removed using the DNA-free™ system included in the kit.
The LCM Staining Kit provides superior staining of frozen sections without compromising RNA integrity to facilitate identification of target cells for LCM prior to isolation with the RNAqueous®-Micro Kit.
Blood Samples: LeukoLOCK™ Total RNA Isolation System
The LeukoLOCK System provides a fast way to isolate nucleic acids from the leukocyte fraction (WBCs) of whole blood. The resulting RNA is inherently depleted of globin mRNA, which improves sample utility for expression profiling and other applications. A modified protocol is used to retain small RNAs.
FFPE Tissues: RecoverAll™ Total Nucleic Acid Isolation Kit for FFPE
The RecoverAll Total Nucleic Acid Isolation Kit extracts total nucleic acid (RNA, including small RNAs, and DNA) from formaldehyde- or paraformaldehyde-fixed, paraffin-embedded (FFPE) tissues. Using this kit, you can now rapidly isolate total RNA containing small RNA from archived tissue samples, enabling retrospective miRNA expression profiling of diseased tissue. Small RNA can then be recovered from the total RNA using the miRNA enrichment protocol and reagents in the mirVana miRNA Isolation Kit.