Search
Identification of miRNA Profiles in Individual Embryonic Stem Cells
MicroRNAs (miRNAs) are a class of small (~22 nt) noncoding RNA involved in post-transcriptional regulation of gene expression. miRNAs interact with messenger RNA (mRNA) to inhibit translation and in some cases, exert their effect by inducing degradation of their target mRNA. In more and more published research studies, miRNAs have been shown to play critical roles in many cellular and disease processes. Characterizing the expression profile of these small RNAs is a first step toward understanding their role in biological pathways, and may identify biomarkers worthy of additional investigation. This article describes a study of miRNA expression in individual mouse embryonic stem cells that was enabled by an early access version of Applied Biosystems Megaplex™ Primer Pools. These new products are not currently supported as a single-cell miRNA profiling tool, but this research highlights the benefits of their low input requirements, unsurpassed sensitivity, wide dynamic range, minimal sample handling, and ease of use.
Characterizing the miRNA Profile in Embryonic Stem Cells
Stem cell biology is a rapidly growing area of research thought to have massive potential for treatment of human disease. Emphasis has been placed on embryonic stem cells (ESCs) because they are totipotent; in other words, they can differentiate into any of the >220 cell types in the adult body. ESCs are derived from the inner cell mass of blastocyst embryos. Their totipotency can be retained in tissue culture, or ESCs in culture can also be directed to mature into specific cell types when given specific differentiation signals.
There is significant evidence that miRNAs are involved in the regulation of stem cell differentiation. Distinct sets of miRNAs have been found exclusively in ESCs or in differentiated adult tissues [1,2]. This research caught the attention of Drs. William Strauss (University of Colorado-Boulder) and Caifu Chen (Research and Development, Applied Biosystems Inc.) who collaborated to investigate whether miRNA expression profiles (signatures) unique to ESCs could be identified. These profiles could be used to characterize and distinguish totipotent ESCs from pluripotent cell types, such as embryoid body (EB) cells, and from terminally-differentiated somatic cells.
There is significant evidence that miRNAs are involved in the regulation of stem cell differentiation. Distinct sets of miRNAs have been found exclusively in ESCs or in differentiated adult tissues [1,2]. This research caught the attention of Drs. William Strauss (University of Colorado-Boulder) and Caifu Chen (Research and Development, Applied Biosystems Inc.) who collaborated to investigate whether miRNA expression profiles (signatures) unique to ESCs could be identified. These profiles could be used to characterize and distinguish totipotent ESCs from pluripotent cell types, such as embryoid body (EB) cells, and from terminally-differentiated somatic cells.
New Megaplex™ Primer Pools Enable miRNA Profiling From Single Cells
To obtain a true miRNA profile from cells at various stages of differentiation, the researchers decided that analyzing individual cells would be the best approach. Clearly this represented a significant technical challenge, but this was considered less important than the biological limitations imposed by the difficulty in culturing ESCs, their tendency to spontaneously differentiate in the absence of feeder cells, and the inherent heterogeneity among EB cells. Dr. Chen recognized this challenge as an opportunity to push the limits of technology under development at Applied Biosystems for a set of miRNA profiling tools that require only very small amounts of RNA for reliable, reproducible, quantitative PCR (see sidebar, "Streamlined miRNA Profiling Using Megaplex™ Primer Pools and TaqMan® MicroRNA Arrays").
70 randomly selected individual cells from a mouse ESC line, day 3 EB cells, and several somatic cell types including 3T3 cells (a well-characterized transformed fibroblast cell line) and primary splenocytes were chosen for miRNA profiling. The cells were lysed by heating to 95°C for 5 minutes in 1X PBS. An early access version of Megaplex™ RT Primers—a pool of 237 stem-loop RT primers from the associated TaqMan MicroRNA Assays—was used to prime reverse transcription of lysates using the TaqMan MicroRNA RT Kit. Since the real-time PCR quantitation step required distributing this minute amount of starting material across 237 independent reactions, it was decided to incorporate a preamplification step. Preamplification preserves the distribution of miRNAs in the starting sample, in particular those miRNAs present at low copy number that would otherwise be diluted out when material was distributed to each independent real-time PCR assay, thereby maintaining overall assay sensitivity. The cDNA was preamplified using a limited number of PCR cycles and an early access version of Megaplex™ PreAmp Primers—a pool of PCR primers from the same 237 TaqMan MicroRNA Assays. Then the preamplification products were subjected to real-time PCR using the 237 individual TaqMan MicroRNA Assays. This workflow enabled single-cell analysis of the expression profiles of most of the murine miRNAs known at the time (miRBase v10, [3]). At the same time, expression data were collected for positive and negative controls, as well as for 21 mRNAs corresponding to transcripts recognized by the stem cell community for their association with ESC differentiation and self-renewal.
70 randomly selected individual cells from a mouse ESC line, day 3 EB cells, and several somatic cell types including 3T3 cells (a well-characterized transformed fibroblast cell line) and primary splenocytes were chosen for miRNA profiling. The cells were lysed by heating to 95°C for 5 minutes in 1X PBS. An early access version of Megaplex™ RT Primers—a pool of 237 stem-loop RT primers from the associated TaqMan MicroRNA Assays—was used to prime reverse transcription of lysates using the TaqMan MicroRNA RT Kit. Since the real-time PCR quantitation step required distributing this minute amount of starting material across 237 independent reactions, it was decided to incorporate a preamplification step. Preamplification preserves the distribution of miRNAs in the starting sample, in particular those miRNAs present at low copy number that would otherwise be diluted out when material was distributed to each independent real-time PCR assay, thereby maintaining overall assay sensitivity. The cDNA was preamplified using a limited number of PCR cycles and an early access version of Megaplex™ PreAmp Primers—a pool of PCR primers from the same 237 TaqMan MicroRNA Assays. Then the preamplification products were subjected to real-time PCR using the 237 individual TaqMan MicroRNA Assays. This workflow enabled single-cell analysis of the expression profiles of most of the murine miRNAs known at the time (miRBase v10, [3]). At the same time, expression data were collected for positive and negative controls, as well as for 21 mRNAs corresponding to transcripts recognized by the stem cell community for their association with ESC differentiation and self-renewal.
miRNA Levels Increase as Differentiation Proceeds
Expression of ESC miRNA was generally found to be low in comparison to the pluripotent and terminally differentiated cells (Figure 1). As cells differentiated into EB cells, both the total number of miRNAs and their expression levels increased. By day 3, significant differences in the amounts of miRNA were detected in partially differentiated EB cells compared to their totipotent ESC precursors (Figure 2). One of the goals of these experiments was to evaluate the variability in miRNA profiles among individual cells in a single ESC line. The data indicated that the cells within a given ESC line represent a mixture of undifferentiated cells and those beginning to differentiate, and that a subset of ~6 miRNAs can be used as markers to distinguish these development stages.
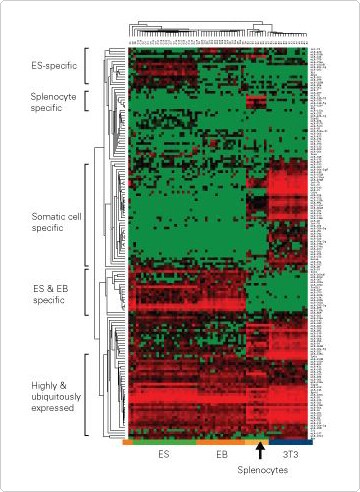
Figure 2. Single Cell Expression Profile of 237 miRNAs and 21 mRNAs in ES, EB, Splenocyte, and 3T3 Cells.

Figure 2. Single Cell Expression Profile of 237 miRNAs and 21 mRNAs in ES, EB, Splenocyte, and 3T3 Cells.
Applied Biosystems tools for miRNA research using TaqMan real-time PCR enabled the analysis of miRNA expression in individual cells. The researchers were able to fulfill their principal goal of identifying miRNA expression signatures for classification of ESCs and differentiated cells, and miRNAs that could be used as biomarkers to define stem cell identity. Because they looked at single cells, they were also able to document significant heterogeneity of miRNA expression among cells at a given stage of differentiation. The coefficient of variance (CV) of cell-to-cell miRNA expression within a population increased significantly from 17% and 18% in 3T3 and ESCs, to 25% and 46% in EBs and hippocampal cells, respectively.
Megaplex Primer Pools and next generation TaqMan MicroRNA Arrays are now available from Applied Biosystems, making it possible for any laboratory to profile miRNA from minute amounts of starting materials. Samples such as tumor biopsies, fluorescently sorted cells, and laser capture microscopy specimens are now much more accessible, but these new tools offer the benefits of simple, rapid processing, and the sensitivity, specificity, and dynamic range of TaqMan real-time PCR to any research program that includes miRNA profiling.
Megaplex Primer Pools and next generation TaqMan MicroRNA Arrays are now available from Applied Biosystems, making it possible for any laboratory to profile miRNA from minute amounts of starting materials. Samples such as tumor biopsies, fluorescently sorted cells, and laser capture microscopy specimens are now much more accessible, but these new tools offer the benefits of simple, rapid processing, and the sensitivity, specificity, and dynamic range of TaqMan real-time PCR to any research program that includes miRNA profiling.
Streamlined miRNA Profiling Using Megaplex™ Primer Pools and TaqMan® MicroRNA ArraysBecause of its high sensitivity, specificity, and broad dynamic range, TaqMan® real-time PCR is considered by many researchers to be the gold standard for gene expression analysis. Likewise, TaqMan MicroRNA Assays have become the technology of choice for quantitative measurement of miRNAs. With a much simpler process and broader dynamic range, Applied Biosystems new Megaplex™ RT Primer Pools and PreAmp Primer Pools for human and rodent species offer an enhanced miRNA profiling workflow that enables experimentation using only minute quantities of input total RNA.
Using Megaplex Primer Pools can substantially reduce sample handling; each set of Megaplex RT Primers contains up to 381 primers for reverse transcription of up to 381 miRNAs simultaneously in a single reaction. For very limited samples, such as those used in the research described here, or when maximum sensitivity is required, an optional preamplification step can be included using Megaplex PreAmp Primers. Although Megaplex Primer Pools can be run with plated individual TaqMan MicroRNA Assays, the ideal profiling workflow utilizes the companion TaqMan MicroRNA Array. The reverse transcribed, preamplified cDNA is simply combined with TaqMan Universal PCR Master Mix, loaded onto an array, and run on the Applied Biosystems 7900HT Fast Real-Time PCR System using universal cycling conditions.
In combination with TaqMan MicroRNA Arrays, Megaplex Primer Pools enable analysis of as little as 1 ng of input total RNA per reaction when run with a preamplification step. As described in this article, this technology can be pushed to significantly lower input amounts although currently RNA input amounts below 1 ng are not fully supported.
For both human and rodent species, comprehensive coverage of the Sanger miRBase v10 database content is achieved through a two TaqMan MicroRNA Array set, each associated with companion Megaplex Primer Pools.
Pairing TaqMan MicroRNA Arrays, Megaplex RT Primers, and PreAmp Primers provides a rapid method for miRNA profiling in human, mouse, or rat samples with minimal sample input requirements. Whether your profiling experiment requires ultimate sensitivity, broad coverage, or both; Megaplex Primer Pools offer the flexibility to reach your research goals.
Using Megaplex Primer Pools can substantially reduce sample handling; each set of Megaplex RT Primers contains up to 381 primers for reverse transcription of up to 381 miRNAs simultaneously in a single reaction. For very limited samples, such as those used in the research described here, or when maximum sensitivity is required, an optional preamplification step can be included using Megaplex PreAmp Primers. Although Megaplex Primer Pools can be run with plated individual TaqMan MicroRNA Assays, the ideal profiling workflow utilizes the companion TaqMan MicroRNA Array. The reverse transcribed, preamplified cDNA is simply combined with TaqMan Universal PCR Master Mix, loaded onto an array, and run on the Applied Biosystems 7900HT Fast Real-Time PCR System using universal cycling conditions.
In combination with TaqMan MicroRNA Arrays, Megaplex Primer Pools enable analysis of as little as 1 ng of input total RNA per reaction when run with a preamplification step. As described in this article, this technology can be pushed to significantly lower input amounts although currently RNA input amounts below 1 ng are not fully supported.
For both human and rodent species, comprehensive coverage of the Sanger miRBase v10 database content is achieved through a two TaqMan MicroRNA Array set, each associated with companion Megaplex Primer Pools.
Pairing TaqMan MicroRNA Arrays, Megaplex RT Primers, and PreAmp Primers provides a rapid method for miRNA profiling in human, mouse, or rat samples with minimal sample input requirements. Whether your profiling experiment requires ultimate sensitivity, broad coverage, or both; Megaplex Primer Pools offer the flexibility to reach your research goals.
References1. Houbaviy HB, Murray MF, Sharp PA (2003) Embryonic stem cell-specific microRNAs. Devel Cell 5: 351–358.
2. Strauss W, Chen C, Lee C-T, Ridzon D (2006) Nonrestrictive developmental regulation of microRNA gene expression. Mamm Genome 17: 833–840.
3. Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36: D154–D158.
2. Strauss W, Chen C, Lee C-T, Ridzon D (2006) Nonrestrictive developmental regulation of microRNA gene expression. Mamm Genome 17: 833–840.
3. Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36: D154–D158.